Difference between revisions of "Template:Article of the week"
Shawndouglas (talk | contribs) (Updated article of the week text.) |
Shawndouglas (talk | contribs) (Updated article of the week text.) |
||
| Line 1: | Line 1: | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 | <div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig1 Easton OJofPubHlthInfo2015 7-3.jpg|240px]]</div> | ||
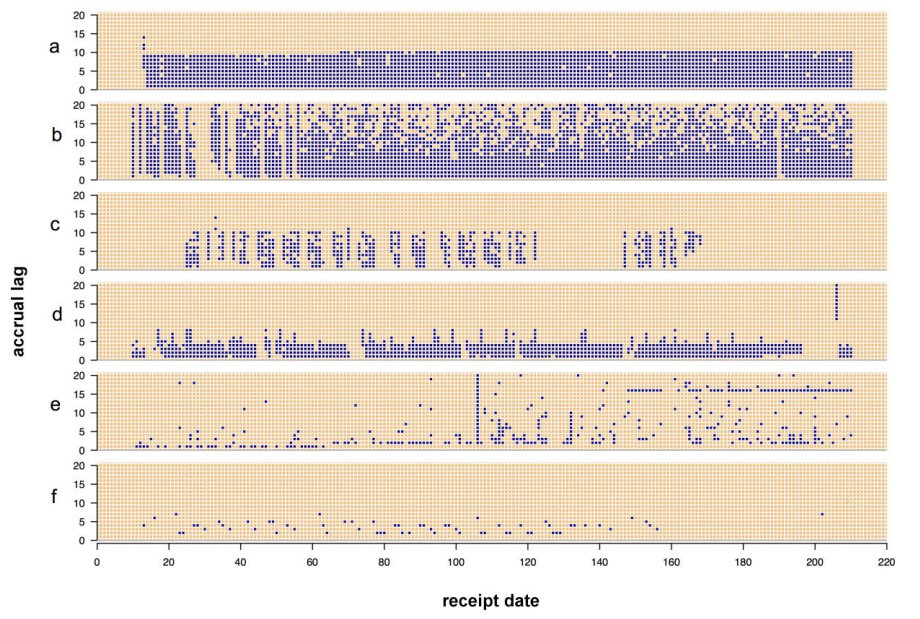
'''"[[Journal: | '''"[[Journal:Visualizing the quality of partially accruing data for use in decision making|Visualizing the quality of partially accruing data for use in decision making]]"''' | ||
Secondary use of clinical health data for near real-time [[Public health laboratory|public health surveillance]] presents challenges surrounding its utility due to data quality issues. Data used for real-time surveillance must be timely, accurate and complete if it is to be useful; if incomplete data are used for surveillance, understanding the structure of the incompleteness is necessary. Such data are commonly aggregated due to privacy concerns. The Distribute project was a near real-time influenza-like-illness (ILI) surveillance system that relied on aggregated secondary clinical health data. The goal of this work is to disseminate the data quality tools developed to gain insight into the data quality problems associated with these data. These tools apply in general to any system where aggregate data are accrued over time and were created through the end-user-as-developer paradigm. Each tool was developed during the exploratory analysis to gain insight into structural aspects of data quality. ('''[[Journal:Visualizing the quality of partially accruing data for use in decision making|Full article...]]''')<br /> | |||
<br /> | <br /> | ||
''Recently featured'': | ''Recently featured'': | ||
: ▪ [[Journal:Digital pathology and anatomic pathology laboratory information system integration to support digital pathology sign-out]] | |||
: ▪ [[Journal:A polyglot approach to bioinformatics data integration: A phylogenetic analysis of HIV-1|A polyglot approach to bioinformatics data integration: A phylogenetic analysis of HIV-1]] | : ▪ [[Journal:A polyglot approach to bioinformatics data integration: A phylogenetic analysis of HIV-1|A polyglot approach to bioinformatics data integration: A phylogenetic analysis of HIV-1]] | ||
: ▪ [[Journal:The systems biology format converter|The systems biology format converter]] | : ▪ [[Journal:The systems biology format converter|The systems biology format converter]] | ||
Revision as of 19:10, 18 July 2016
"Visualizing the quality of partially accruing data for use in decision making"
Secondary use of clinical health data for near real-time public health surveillance presents challenges surrounding its utility due to data quality issues. Data used for real-time surveillance must be timely, accurate and complete if it is to be useful; if incomplete data are used for surveillance, understanding the structure of the incompleteness is necessary. Such data are commonly aggregated due to privacy concerns. The Distribute project was a near real-time influenza-like-illness (ILI) surveillance system that relied on aggregated secondary clinical health data. The goal of this work is to disseminate the data quality tools developed to gain insight into the data quality problems associated with these data. These tools apply in general to any system where aggregate data are accrued over time and were created through the end-user-as-developer paradigm. Each tool was developed during the exploratory analysis to gain insight into structural aspects of data quality. (Full article...)
Recently featured: