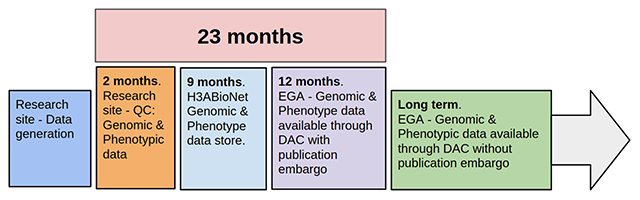
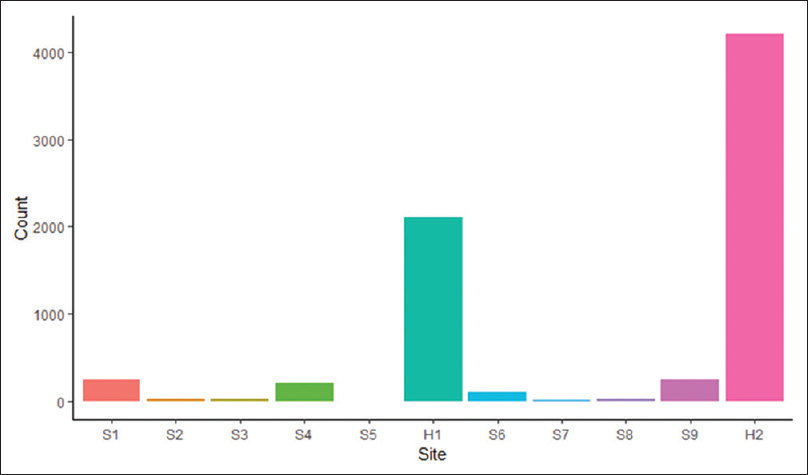
Featured article of the week: December 23–31:"Building infrastructure for African human genomic data management"
Human genomic data are large and complex, and require adequate infrastructure for secure storage and transfer. The National Institutes of Health (NIH) and The Wellcome Trust have funded multiple projects on genomic research, including the Human Heredity and Health in Africa (H3Africa) initiative, and data are required to be deposited into the public domain. The European Genome-phenome Archive (EGA) is a repository for sequence and genotype data where data access is controlled by access committees. Access is determined by a formal application procedure for the purpose of secure storage and distribution, which must be in line with the informed consent of the study participants. H3Africa researchers based in Africa and generating their own data can benefit tremendously from the data sharing capabilities of the internet by using the appropriate technologies. The H3Africa Data Archive is an effort between the H3Africa data generating projects, H3ABioNet, and the EGA to store and submit genomic data to public repositories. (Full article...)
Featured article of the week: December 16–22:
"Process variation detection using missing data in a multihospital community practice anatomic pathology laboratory"
Barcode-driven workflows reduce patient identification errors. Missing process timestamp data frequently confound our health system's pending lists and appear as actions left undone. Anecdotally, it was noted that missing data could be found when there is procedure noncompliance. This project was developed to determine if missing timestamp data in the histology barcode-driven workflow correlated with other process variations, procedure noncompliance, or is an indicator of workflows needing focus for improvement projects. Data extracts of timestamp data from January 1, 2018 to December 15, 2018 for the major histology process steps were analyzed for missing data. Case-level analysis to determine the presence or absence of expected barcoding events was performed on 1031 surgical pathology cases to determine the cause of the missing data and determine if additional data variations or procedure noncompliance events were present. The data variations were classified according to a scheme defined in the study. (Full article...)
|
Featured article of the week: December 9–15:
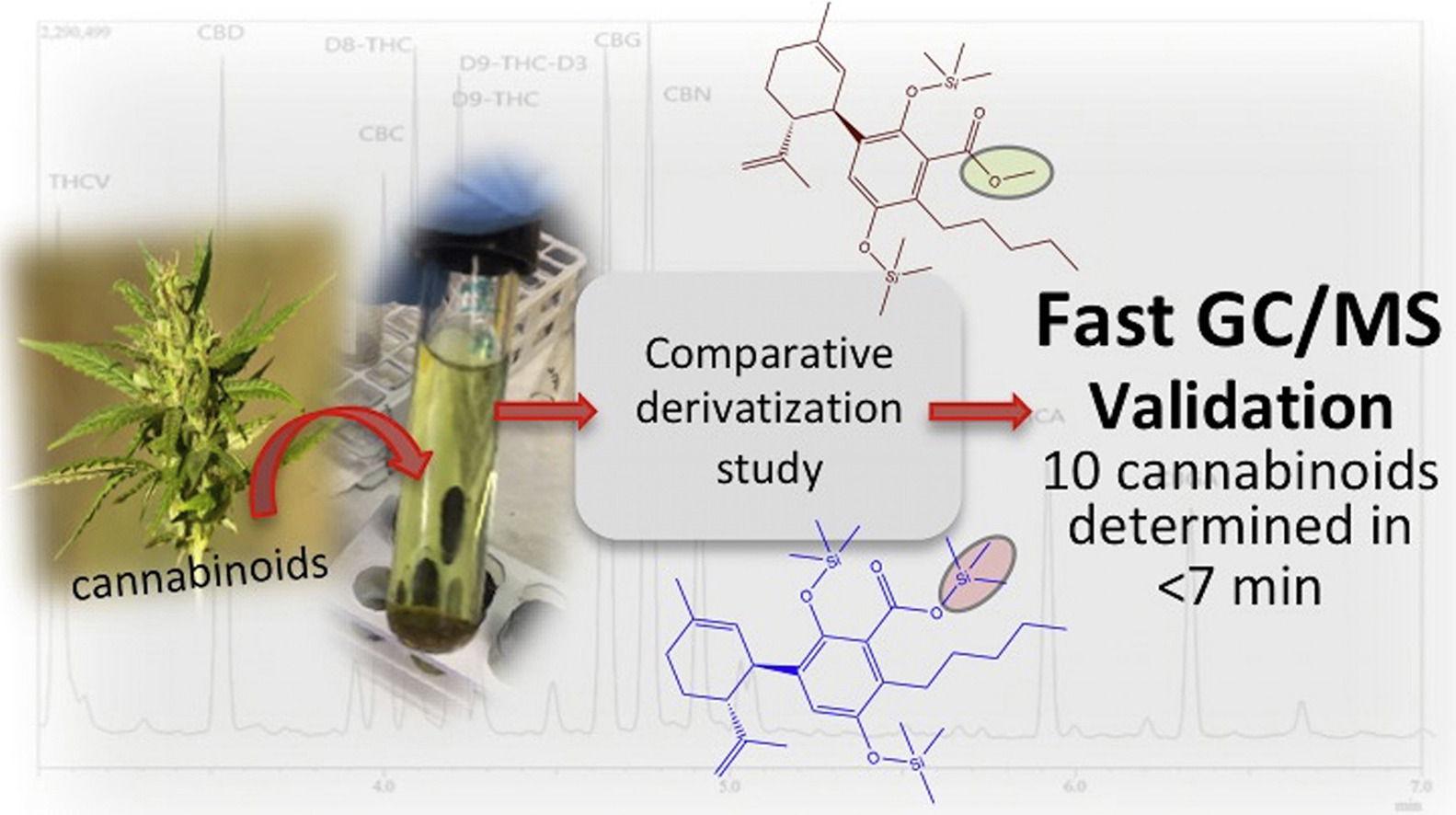
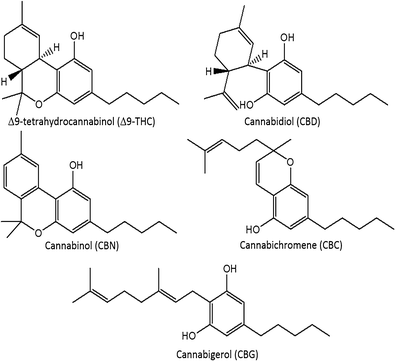
"Development and validation of a fast gas chromatography–mass spectrometry method for the determination of cannabinoids in Cannabis sativa L"
A routine method for determining cannabinoids in Cannabis sativa L. inflorescence, based on fast gas chromatography coupled to mass spectrometry (fast GC-MS), was developed and validated. To avoid the decarboxylation of the carboxyl group of cannabinoids, different derivatization approaches—i.e., silylation and esterification (diazomethane-mediated) reagents and solvents (pyridine or ethyl acetate)—were tested. The methylation significantly increased the signal-to-noise ratio of all carboxylic cannabinoids, except for cannabigerolic acid (CBGA). Since diazomethane is not commercially available, is considered a hazardous reactive, and requires one-day synthesis by specialized chemical staff, the process of silylation was used along the entire validation of a routine method. The method gave a fast (total analysis time < 7.0 min) and satisfactory resolution (R > 1.1), with a good repeatability (intraday < 8.38%; interday < 11.10%) and sensitivity (LOD < 11.20 ng/mL). (Full article...)
|
Featured article of the week: December 2–8:
"Design and refinement of a data quality assessment workflow for a large pediatric research network"
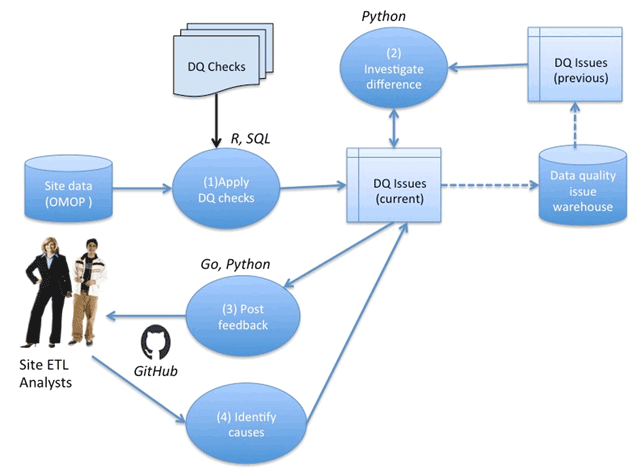
Clinical data research networks (CDRNs) aggregate electronic health record (EHR) data from multiple hospitals to enable large-scale research. A critical operation toward building a CDRN is conducting continual evaluations to optimize data quality. The key challenges include determining the assessment coverage on big datasets, handling data variability over time, and facilitating communication with data teams. This study presents the evolution of a systematic workflow for data quality assessment in CDRNs.
Using a specific CDRN as a use case, a workflow was iteratively developed and packaged into a toolkit. The resultant toolkit comprises 685 data quality checks to identify any data quality issues, procedures to reconciliate with a history of known issues, and a contemporary GitHub-based reporting mechanism for organized tracking.
During the first two years of network development, the toolkit assisted in discovering over 800 data characteristics and resolving over 1400 programming errors. Longitudinal analysis indicated that the variability in time to resolution (15day mean, 24day IQR) is due to the underlying cause of the issue, perceived importance of the domain, and the complexity of assessment. (Full article...)
|
Featured article of the week: November 25–December 1:
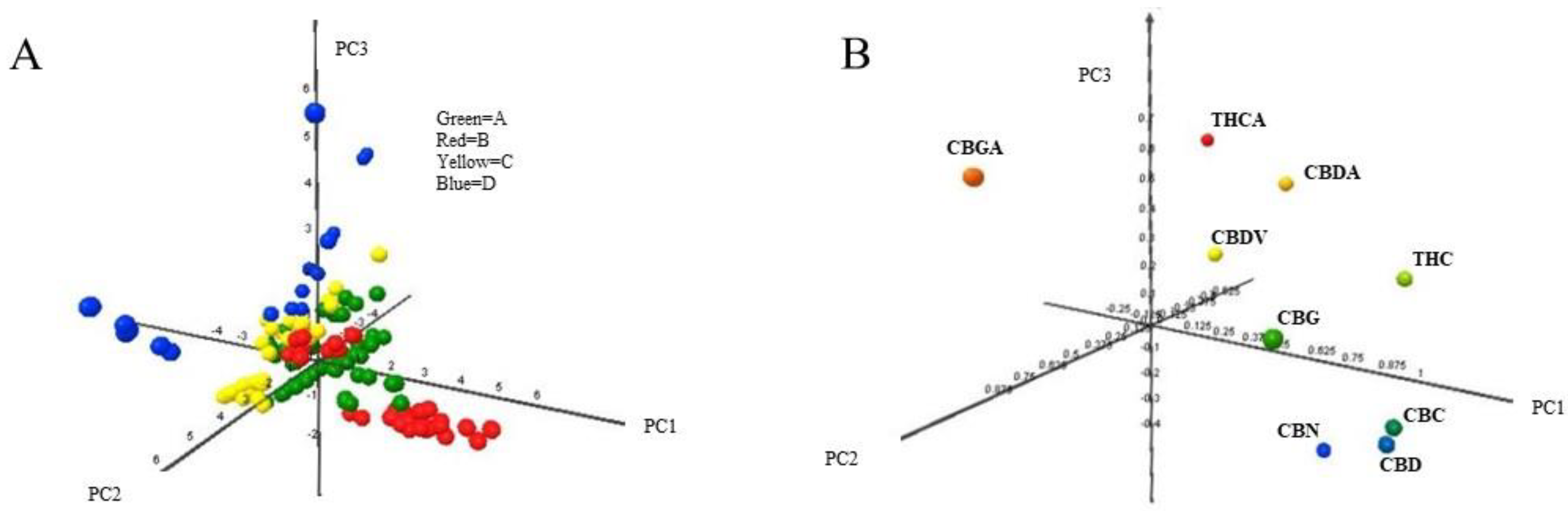
"Identification of Cannabis sativa L. (hemp) retailers by means of multivariate analysis of cannabinoids"
In this work, the concentration of nine cannabinoids—six neutral cannabinoids (THC, CBD, CBC, CBG, CBN, and CBDV) and three acidic cannabinoids (THCA, CBGA, and CBDA)—was used to identify the Italian retailers of Cannabis sativa L. (hemp), reinforcing the idea that the practice of categorizing hemp samples only using THC and CBD is inadequate. A high-performance liquid chromatography–tandem mass spectrometry (HPLC-MS/MS) method was developed for screening and simultaneously analyzing the nine cannabinoids in 161 hemp samples sold by four retailers located in different Italian cities. The hemp samples dataset was analyzed by univariate and multivariate analysis, with the aim to identify the associated hemp retailers without using any other information on the hemp samples such as Cannabis strains, seeds, soil and cultivation characteristics, geographical origin, product storage, etc. The univariate analysis highlighted that the hemp samples could not be differentiated by using any of the nine cannabinoids analyzed. (Full article...)
|
Featured article of the week: November 18–24:
"Data sharing at scale: A heuristic for affirming data cultures"
Addressing the most pressing contemporary social, environmental, and technological challenges will require integrating insights and sharing data across disciplines, geographies, and cultures. Strengthening international data sharing networks will not only demand advancing technical, legal, and logistical infrastructure for publishing data in open, accessible formats; it will also require recognizing, respecting, and learning to work across diverse data cultures. This essay introduces a heuristic for pursuing richer characterizations of the “data cultures” at play in international, interdisciplinary data sharing. The heuristic prompts cultural analysts to query the contexts of data sharing for a particular discipline, institution, geography, or project at seven scales: the meta, macro, meso, micro, techno, data, and nano. The essay articulates examples of the diverse cultural forces acting upon and interacting with researchers in different communities at each scale. The heuristic we introduce in this essay aims to elicit from researchers the beliefs, values, practices, incentives, and restrictions that impact how they think about and approach data sharing. Rather than represent an effort to iron out differences between disciplines, this essay instead intends to showcase and affirm the diversity of traditions and modes of analysis that have shaped how data gets collected, organized, and interpreted in diverse settings. (Full article...)
|
Featured article of the week: November 11–17:
"Design and evaluation of a LIS-based autoverification system for coagulation assays in a core clinical laboratory"
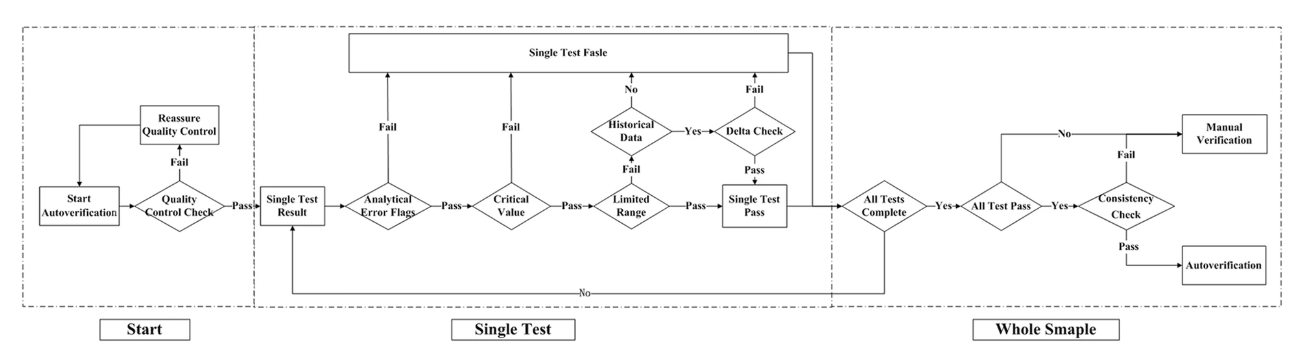
n autoverification system for coagulation consists of a series of rules that allows normal data to be released without manual verification. With new advances in medical informatics, the laboratory information system (LIS) has growing potential for the use of autoverification, allowing rapid and accurate verification of clinical laboratory tests. The purpose of the study is to develop and evaluate a LIS-based autoverification system for validation and efficiency.
Autoverification decision rules—including quality control, analytical error flag, critical value, limited range check, delta check, and logical check rules, as well as patient’s historical information—were integrated into the LIS. Autoverification limit ranges was constructed based on 5% and 95% percentiles. The four most commonly used coagulation assays—prothrombin time (PT), activated partial thromboplastin time (APTT), thrombin time (TT), and fibrinogen (FBG)—were followed by the autoverification protocols. (Full article...)
|
Featured article of the week: November 4–10:
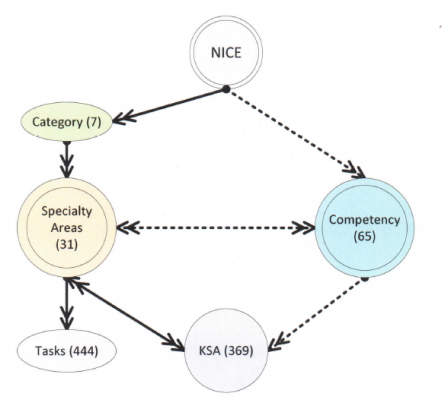
"CyberMaster: An expert system to guide the development of cybersecurity curricula"
The growing number of reported cyberattacks poses a difficult challenge to individuals, governments, and organizations. Adequate protection of information systems urgently requires a cybersecurity-educated workforce trained using a curriculum that covers the essential skills required for different cybersecurity work roles. The goal of the CyberMaster expert system is to assist inexperienced instructors with cybersecurity course design. It is an intelligent system that uses visual feedback to guide the user through the design process. Initial test executions show the promise of such a system in addressing the enormous shortage of cybersecurity experts currently available for designing courses and training programs. (Full article...)
|
Featured article of the week: October 21–November 3:
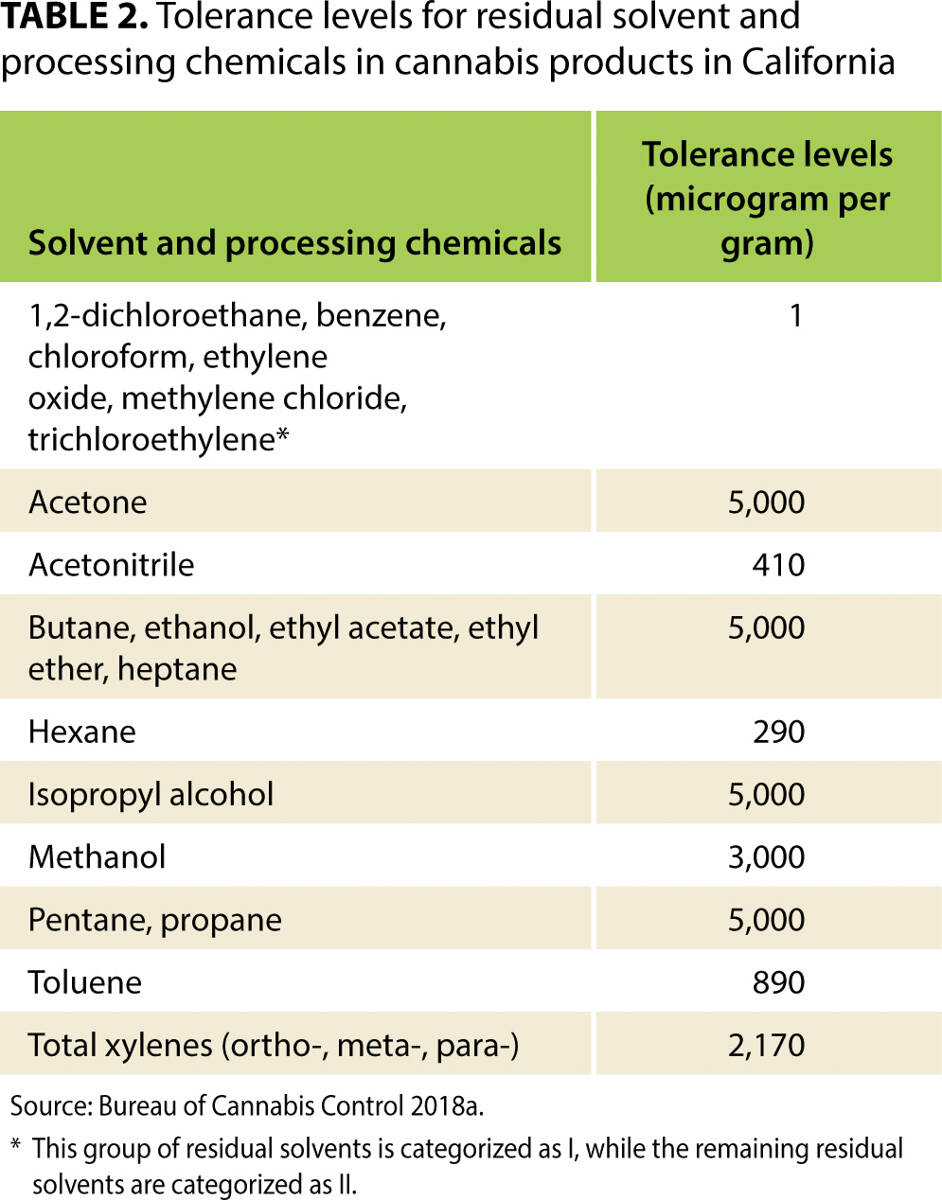
"Costs of mandatory cannabis testing in California"
Every batch of cannabis sold legally in California must be tested for more than 100 contaminants. These contaminants include 66 pesticides, for 21 of which the state's tolerance is zero. For many other substances, tolerance levels are much lower than those allowed for food products in California. This article reviews the state's testing regulations in context—including maximum allowable tolerance levels—and uses primary data collected from California's major cannabis testing laboratories and several cannabis testing equipment manufacturers, as well as a variety of expert opinions, to estimate the cost per pound of testing under the state's framework. We also estimate the cost of collecting samples, which depends on the distance between cannabis distributors and laboratories. We find that, if a batch fails mandatory tests, the value of cannabis that must be destroyed accounts for a large share of total testing costs, more than the cost of the tests that laboratories perform. Findings from this article will help readers understand the effects of California's testing regime on the price of legal cannabis in the state, and understand how testing may add value to products that have passed a series of tests that aim to validate their safety. (Full article...)
|
Featured article of the week: October 14–20:
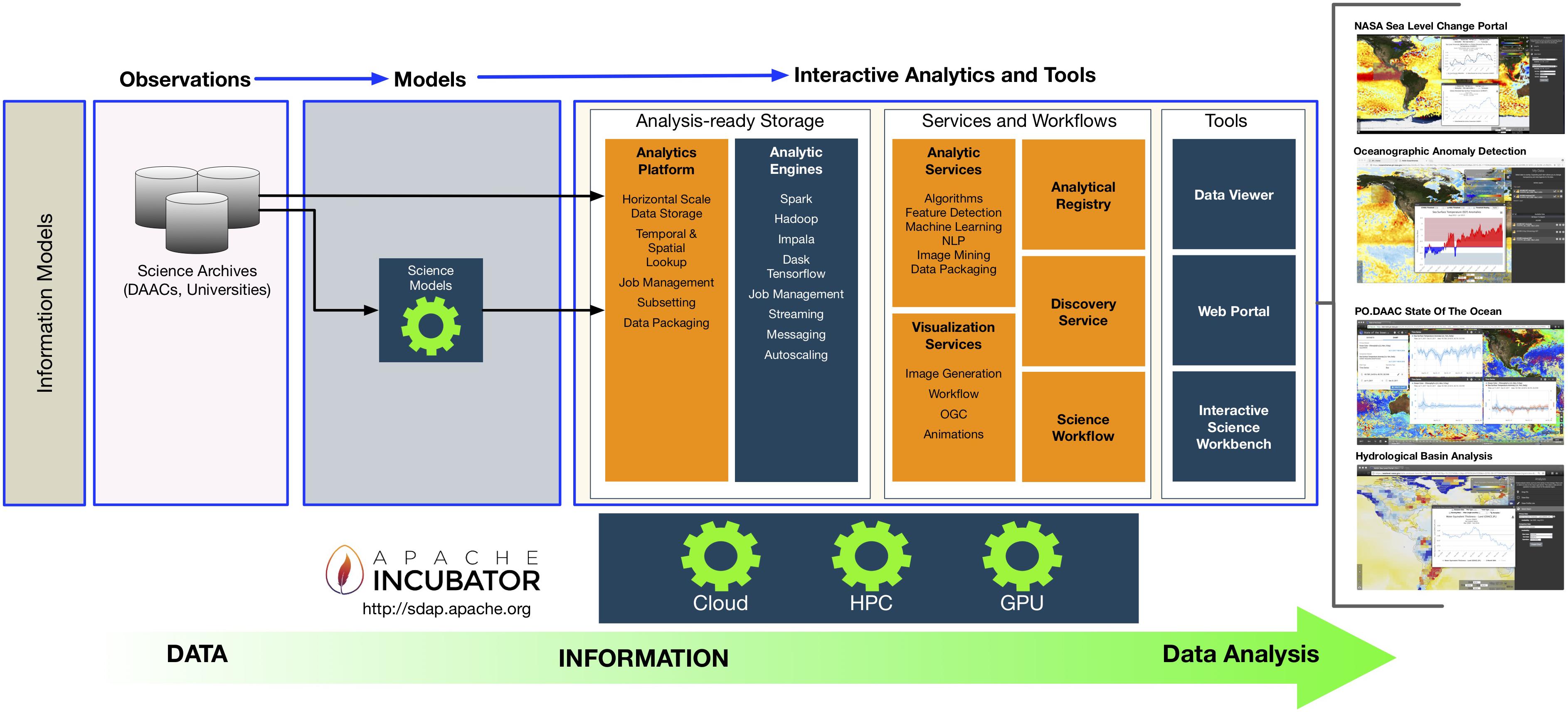
"An integrated data analytics platform"
A scientific integrated data analytics platform (IDAP) is an environment that enables the confluence of resources for scientific investigation. It harmonizes data, tools, and computational resources to enable the research community to focus on the investigation rather than spending time on security, data preparation, management, etc. OceanWorks is a National Aeronautics and Space Administration (NASA) technology integration project to establish a cloud-based integrated ocean science data analytics platform for managing ocean science research data at NASA’s Physical Oceanography Distributed Active Archive Center (PO.DAAC). The platform focuses on advancement and maturity by bringing together several NASA open-source, big data projects for parallel analytics, anomaly detection, in situ-to-satellite data matching, quality-screened data subsetting, search relevancy, and data discovery. (Full article...)
|
Featured article of the week: October 7–13:
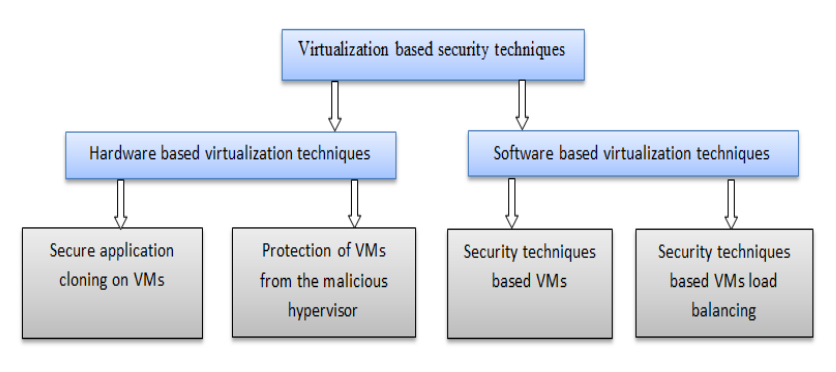
"Virtualization-based security techniques on mobile cloud computing: Research gaps and challenges"
The principle constraints of mobile devices are their limited resources, including processing capability, storage space, and battery life. However, cloud computing offers a means of vast computing resources and services. With it a new idea emerged, the inclusion of cloud computing into mobile devices such as smartphones, tablet, and other personal digital assistants (PDA) to augment their capacities, providing a robust technology called mobile cloud computing (MCC). Although MCC has brought many advantages to mobile users, it also still suffers from the security and privacy issues of data while hosted on virtual machines (VM) on remote cloud’s servers. Currently, the eyes of security experts are turned towards the virtualization-based security techniques used either on the cloud or on mobile devices. The new challenge is to develop secure methods in order to authenticate highly sensitive digital content. (Full article...)
|
Featured article of the week: September 30–October 6:
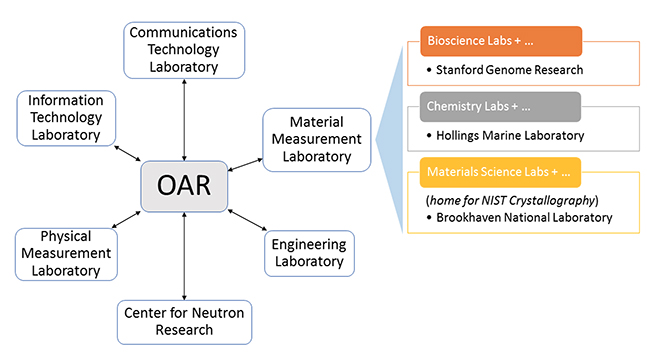
"Building open access to research (OAR) data infrastructure at NIST"
As a National Metrology Institute (NMI), the U.S. National Institute of Standards and Technology (NIST) scientists, engineers, and technology experts conduct research across a full spectrum of physical science domains. NIST is a non-regulatory agency within the U.S. Department of Commerce with a mission to promote U.S. innovation and industrial competitiveness by advancing measurement science, standards, and technology in ways that enhance economic security and improve our quality of life. NIST research results in the production and distribution of standard reference materials, [[calibration services, and datasets. These are generated from a wide range of complex laboratory instrumentation, expert analyses, and calibration processes. In response to a government open data policy, and in collaboration with the broader research community, NIST has developed a federated Open Access to Research (OAR) scientific data infrastructure aligned with FAIR (findable, accessible, interoperable, reusable) data principles. (Full article...)
|
Featured article of the week: September 23–29:
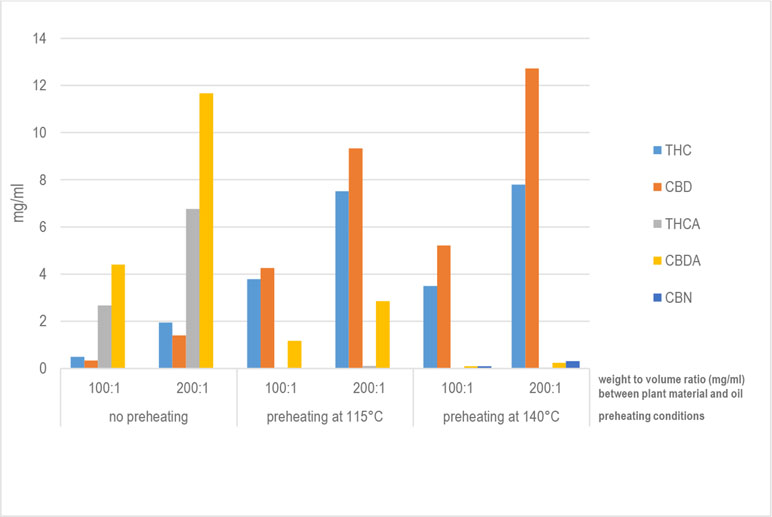
"Development of standard operating protocols for the optimization of Cannabis-based formulations for medical purposes"
Under current legislation in Italy, using the Cannabis plant for medical purposes requires administering it orally in the form of a decoction or as Cannabis oil extract. The scientific literature reports a number of preparation methods, mainly for oils, but no study is available that compares thoroughly, from a technological viewpoint, the Cannabis-based formulations currently administered to patients. With this in mind, this research work aimed to carry out specific formulation studies to design standard operating procedures for the preparation and optimization of Cannabis-based galenic formulations. Both decoctions and oils were prepared under different operating conditions to identify the most efficient process for the production of formulations with a high concentration of decarboxylated delta-9-tetrahydrocannabinol (THC) and cannabidiol (CBD). Regarding Cannabis oil, a new procedure has been developed that allows significantly higher recovery rates for THC and CBD compared with those for water-based extraction methods (decoction) and those for oil-based methods currently in use. (Full article...)
|
Featured article of the week: September 16–22:
"Next steps for access to safe, secure DNA synthesis"
The DNA synthesis industry has, since the invention of gene-length synthesis, worked proactively to ensure synthesis is carried out securely and safely. Informed by guidance from the U.S. government, several of these companies have collaborated over the last decade to produce a set of best practices for customer and sequence screening prior to manufacture. Taken together, these practices ensure that synthetic DNA is used to advance research that is designed and intended for public benefit. With increasing scale in the industry and expanding capability in the synthetic biology toolset, it is worth revisiting current practices to evaluate additional measures to ensure the continued safety and wide availability of DNA synthesis. Here we encourage specific steps, in part derived from successes in the cybersecurity community, that can ensure synthesis screening systems stay well ahead of emerging challenges, to continue to enable responsible research advances. Gene synthesis companies, science and technology funders, policymakers, and the scientific community as a whole have a shared duty to continue to minimize risk and maximize the safety and security of DNA synthesis to further power world-changing developments in advanced biological manufacturing, agriculture, drug development, healthcare, and energy. (Full article...)
|
Featured article of the week: September 9–15:
"Smart grids and ethics: A case study"
This case study explores the principal ethical issues that occur in the use of smart information systems (SIS) in smart grids and offers suggestions as to how they might be addressed. Key issues highlighted in the literature are reviewed. The empirical case study describes one of the largest distribution system operators (DSOs) in the Netherlands. The aim of this case study is to identify which ethical issues arise from the use of SIS in smart grids, the current efforts of the organization to address them, and whether practitioners are facing additional issues not addressed in current literature. The literature review highlights mainly ethical issues around health and safety, privacy and informed consent, cyber-risks and energy security, affordability, equity, and sustainability. The key topics raised by interviewees revolved around privacy and to some extent cybersecurity. (Full article...)
|
Featured article of the week: September 2–8:
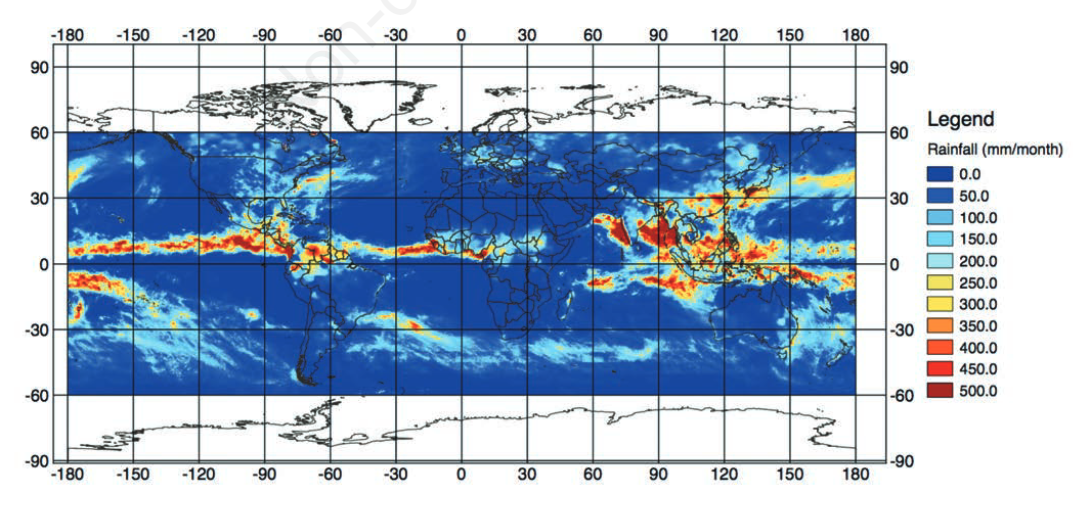
"Japan Aerospace Exploration Agency’s public-health monitoring and analysis platform: A satellite-derived environmental information system supporting epidemiological study"
Since the 1970s, Earth-observing satellites collect increasingly detailed environmental information on land cover, meteorological conditions, environmental variables, and air pollutants. This information spans the entire globe, and its acquisition plays an important role in epidemiological analysis when in situ data are unavailable or spatially and/or temporally sparse. In this paper, we present the development of the Japan Aerospace Exploration Agency’s (JAXA) Public-health Monitoring and Analysis Platform, a user-friendly, web-based system providing environmental data on shortwave radiation, rainfall, soil moisture, the normalized difference vegetation index, aerosol optical thickness, land surface temperature and altitude. (Full article...)
|
Featured article of the week: August 26–September 1:
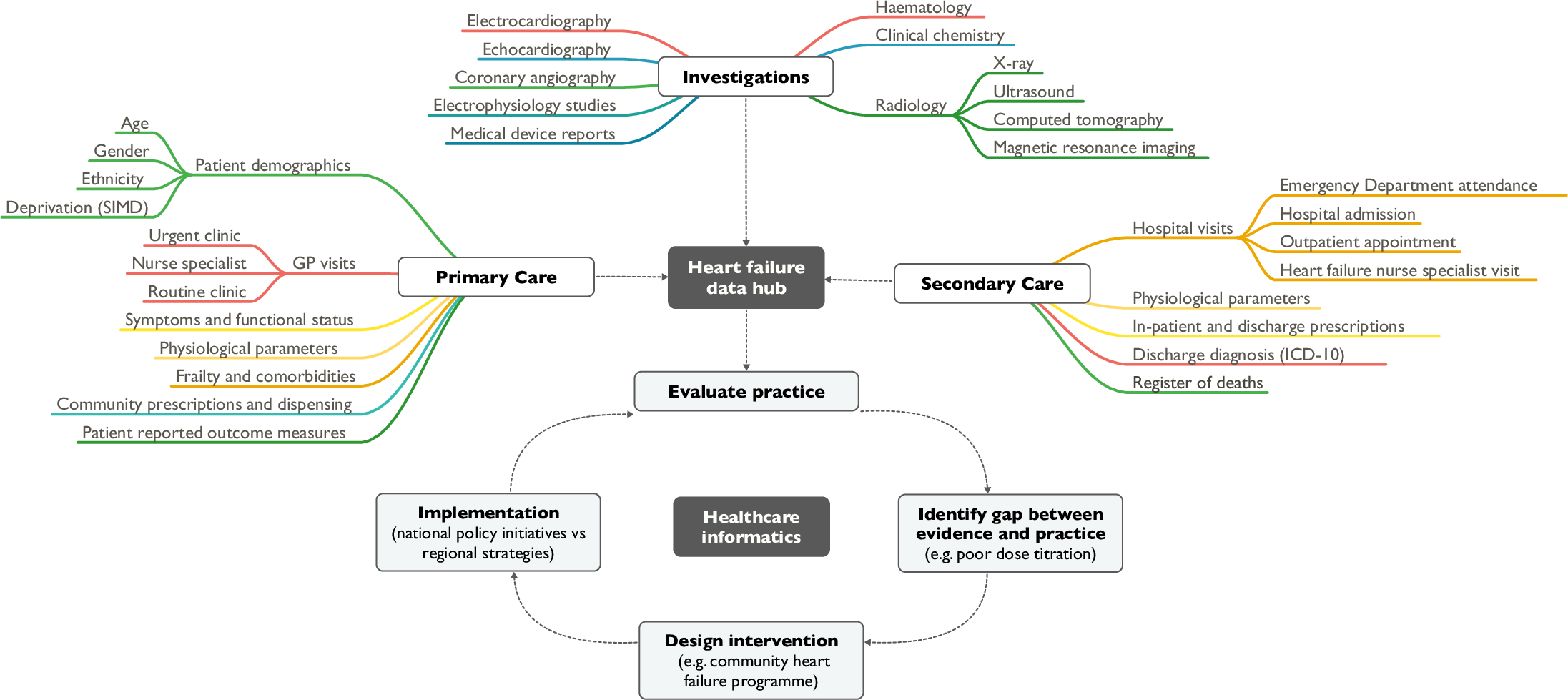
"Heart failure and healthcare informatics"
As biomedical research expands our armory of effective, evidence-based therapies, there is a corresponding need for high-quality implementation science—the study of strategies to integrate and embed research advances into clinical practice. Large-scale collection and analysis of routinely collected healthcare data may facilitate this in three main ways. Firstly, evaluation of key healthcare metrics can help to identify the areas of practice that differ most from guideline recommendations. Secondly, with sufficiently granular data, it may be possible to detect the underlying drivers of deficiencies in practice. Thirdly, longitudinal data collection should enable us to evaluate large-scale policy initiatives and compare the effectiveness of differing strategies on process and patient outcomes. (Full article...)
|
Featured article of the week: August 19–25:
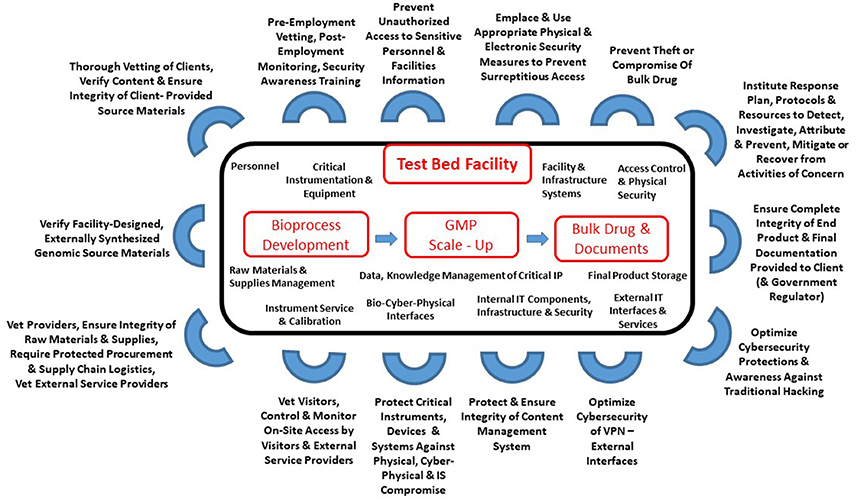
"Cyberbiosecurity for biopharmaceutical products"
Cyberbiosecurity is an emerging discipline that addresses the unique vulnerabilities and threats that occur at the intersection of cyberspace and biotechnology. Advances in technology and manufacturing are increasing the relevance of cyberbiosecurity to the biopharmaceutical manufacturing community in the United States. Threats may be associated with the biopharmaceutical product itself or with the digital thread of manufacturing of biopharmaceuticals, including those that relate to supply chain and cyberphysical systems. Here, we offer an initial examination of these cyberbiosecurity threats as they stand today, as well as introductory steps toward paths for mitigation of cyberbiosecurity risk for a safer, more secure future. (Full article...)
|
Featured article of the week: August 12–18:
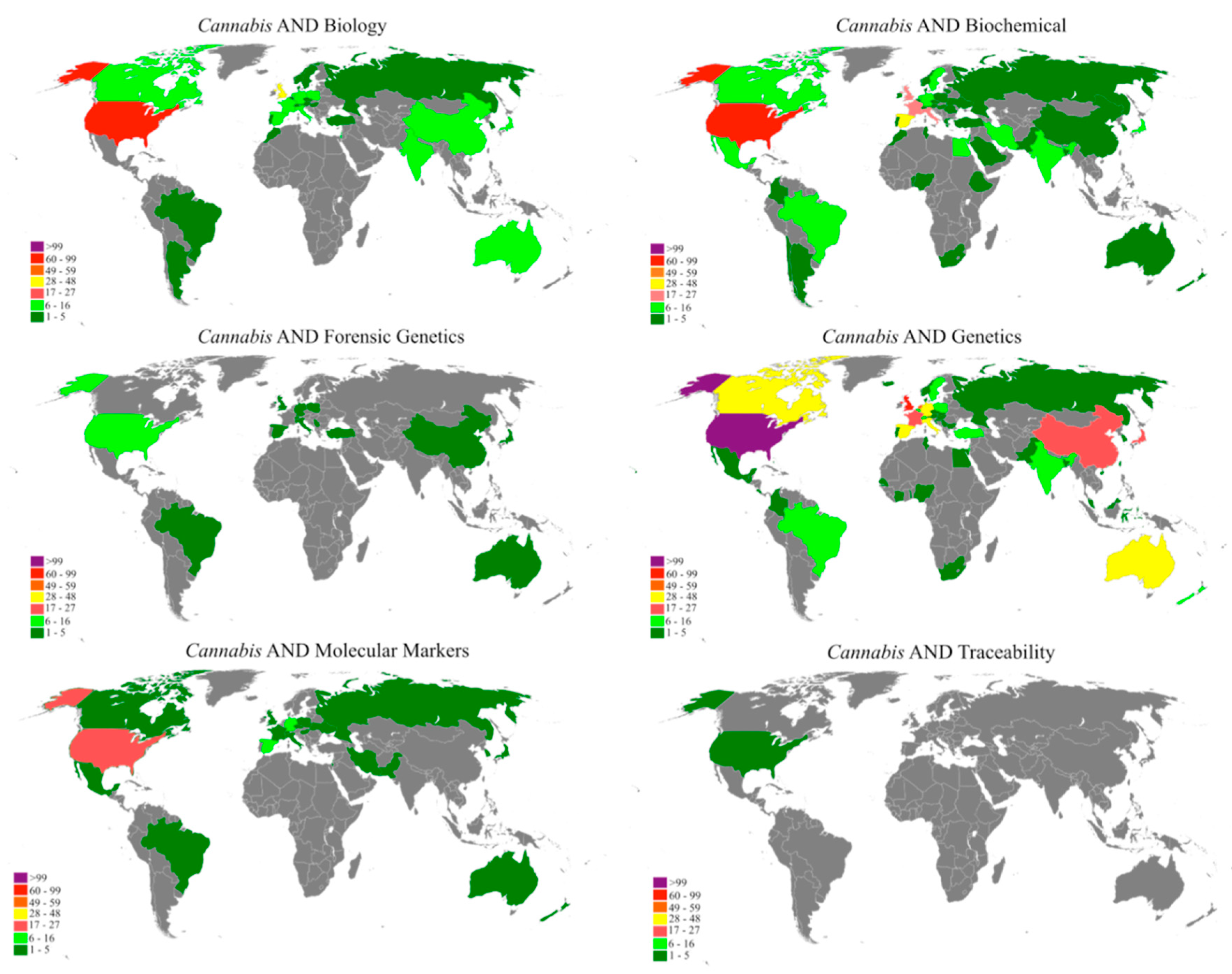
"A bibliometric analysis of Cannabis publications: Six decades of research and a gap on studies with the plant"
In this study we performed a bibliometric analysis focusing on the general patterns of scientific publications about Cannabis, revealing their trends and limitations. Publications related to Cannabis, released from 1960 to 2017, were retrieved from the Scopus database using six search terms. The search term “Genetics” returned 53.4% of publications, while “forensic genetics” and “traceability” represented 2.3% and 0.1% of the publications, respectively. However, 43.1% of the studies were not directly related to Cannabis and, in some cases, Cannabis was just used as an example in the text. A significant increase in publications was observed after 2001, with most of the publications coming from Europe, followed by North America. Although the term "Cannabis" was found in the title, abstract, or keywords of 1284 publications, we detected a historical gap in studies on the plant. (Full article...)
|
Featured article of the week: August 5–11:
"Leaner and greener analysis of cannabinoids"
There is an explosion in the number of labs analyzing cannabinoids in marijuana (Cannabis sativa L., Cannabaceae); however, existing methods are inefficient, require expert analysts, and use large volumes of potentially environmentally damaging solvents. The objective of this work was to develop and validate an accurate method for analyzing cannabinoids in cannabis raw materials and finished products that is more efficient and uses fewer toxic solvents. A method using high-performance liquid chromatography (HPLC) with diode-array detection (DAD) was developed for eight cannabinoids in Cannabis flowers and oils using a statistically guided optimization plan based on the principles of green chemistry. A single-laboratory validation determined the linearity, selectivity, accuracy, repeatability, intermediate precision, limit of detection, and limit of quantitation of the method. Amounts of individual cannabinoids above the limit of quantitation in the flowers ranged from 0.02 to 14.9% concentration (w/w), with repeatability ranging from 0.78 to 10.08% relative standard deviation. (Full article...)
|
Featured article of the week: July 29–August 4:
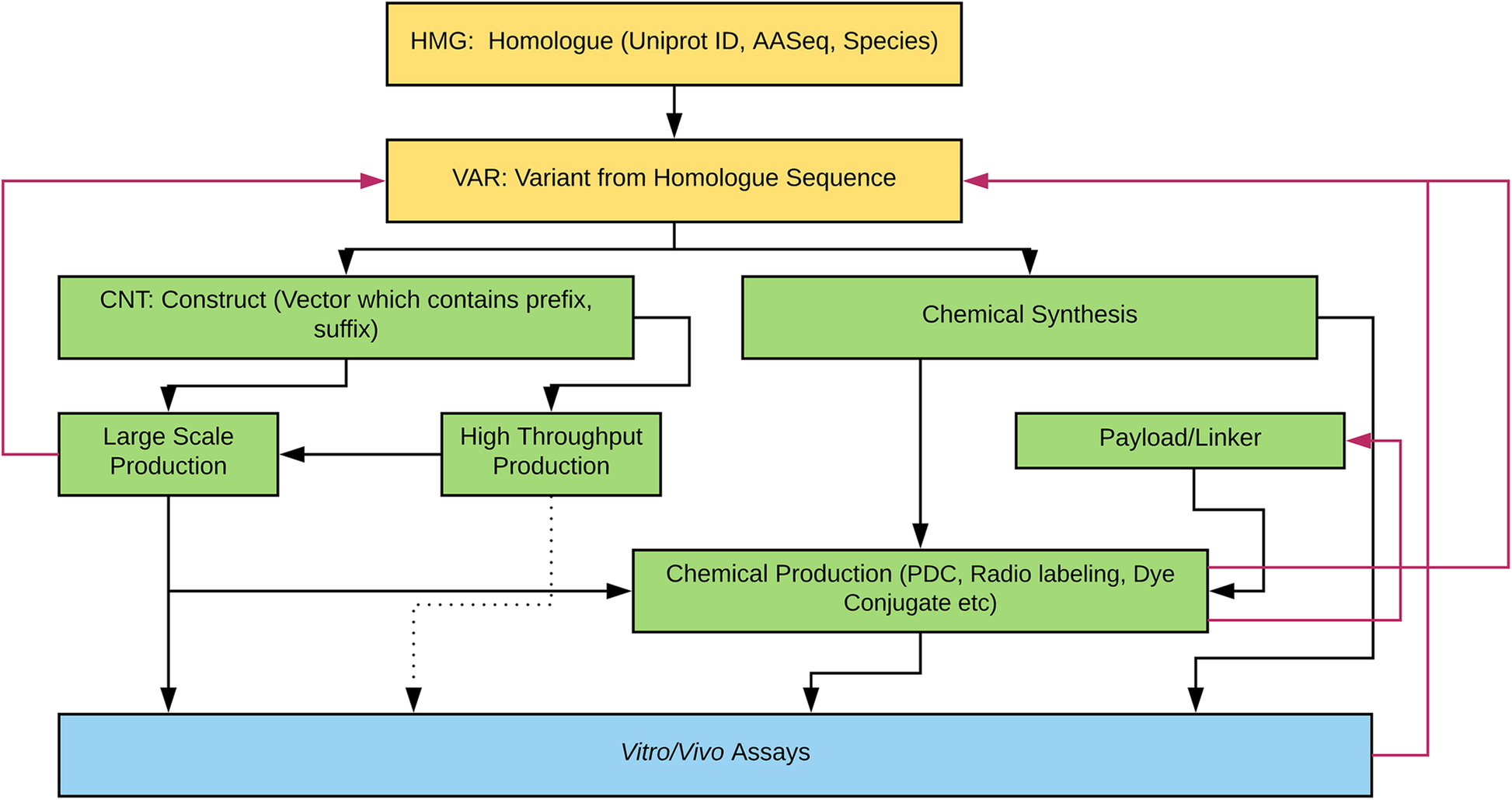
"Laboratory information management software for engineered mini-protein therapeutic workflow"
Protein-based therapeutics are one of the fastest growing classes of novel medical interventions in areas such as cancer, infectious disease, and inflammation. Protein engineering plays an important role in the optimization of desired therapeutic properties such as reducing immunogenicity, increasing stability for storage, increasing target specificity, etc. One category of protein therapeutics is nature-inspired bioengineered cystine-dense peptides (CDPs) for various biological targets. These engineered proteins are often further modified by synthetic chemistry. For example, candidate mini-proteins can be conjugated into active small molecule drugs. We refer to modified mini-proteins as "optides" (optimized peptides). To efficiently serve the multidisciplinary lab scientists with varied therapeutic portfolio research goals in a non-commercial setting, a cost-effective, extendable laboratory information management system (LIMS) is/was needed. (Full article...)
|
Featured article of the week: July 22–28:
"Defending our public biological databases as a global critical infrastructure"
Progress in modern biology is being driven, in part, by the large amounts of freely available data in public resources such as the International Nucleotide Sequence Database Collaboration (INSDC), the world's primary database of biological sequence (and related) information. INSDC and similar databases have dramatically increased the pace of fundamental biological discovery and enabled a host of innovative therapeutic, diagnostic, and forensic applications. However, as high-value, openly shared resources with a high degree of assumed trust, these repositories share compelling similarities to the early days of the internet. Consequently, as public biological databases continue to increase in size and importance, we expect that they will face the same threats as undefended cyberspace. There is a unique opportunity, before a significant breach and loss of trust occurs, to ensure they evolve with quality and security as a design philosophy rather than costly “retrofitted” mitigations. This perspective article surveys some potential quality assurance and security weaknesses in existing open genomic and proteomic repositories, describes methods to mitigate the likelihood of both intentional and unintentional errors, and offers recommendations for risk mitigation based on lessons learned from cybersecurity. (Full article...)
|
Featured article of the week: July 15–21:
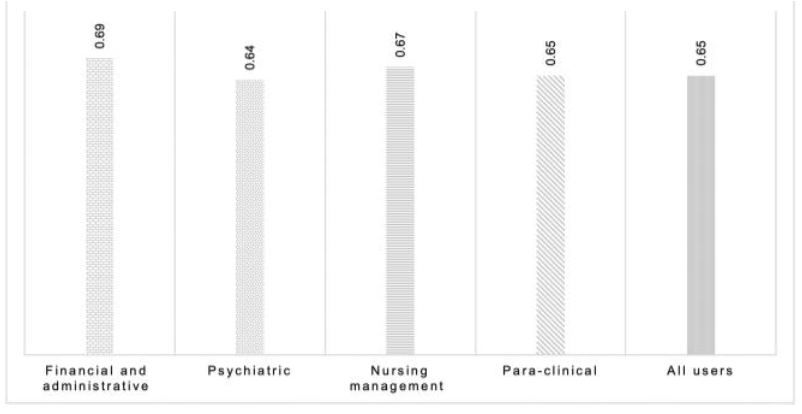
"Determining the hospital information system (HIS) success rate: Development of a new instrument and case study"
A hospital information system (HIS) is a type of health information system which is widely used in clinical settings. Determining the success rate of a HIS is an ongoing area of research since its implications are of interest for researchers, physicians, and managers. In the present study, we develop a novel instrument to measure HIS success rate based on users’ viewpoints in a teaching hospital. The study was conducted in Ibn-e Sina and Dr. Hejazi Psychiatry Hospital and education center in Mashhad, Iran. The instrument for data collection was a self-administered structured questionnaire based on the information systems success model (ISSM), covering seven dimensions, which includes system quality, information quality, service quality, system use, usefulness, satisfaction, and net benefits. The verification of content validity was carried out by an expert panel. The internal consistency of dimensions was measured by Cronbach’s alpha. Pearson’s correlation coefficient was calculated to evaluate the significance of associations between dimensions. The HIS success rate on users’ viewpoints was determined. (Full article...)
|
Featured article of the week: July 08–14:
"Smart information systems in cybersecurity: An ethical analysis"
This report provides an overview of the current implementation of smart information systems (SIS) in the field of cybersecurity. It also identifies the positive and negative aspects of using SIS in cybersecurity, including ethical issues which could arise while using SIS in this area. One company working in the industry of telecommunications (Company A) is analysed in this report. Further specific ethical issues that arise when using SIS technologies in Company A are critically evaluated. Finally, conclusions are drawn on the case study, and areas for improvement are suggested. Increasing numbers of items are becoming connected to the internet. Cisco—a global leader in information technology, networking, and cybersecurity—estimates that more than 8.7 billion devices were connected to the internet by the end of 2012, a number that will likely rise to over 40 billion in 2020. (Full article...)
|
Featured article of the week: July 01–07:
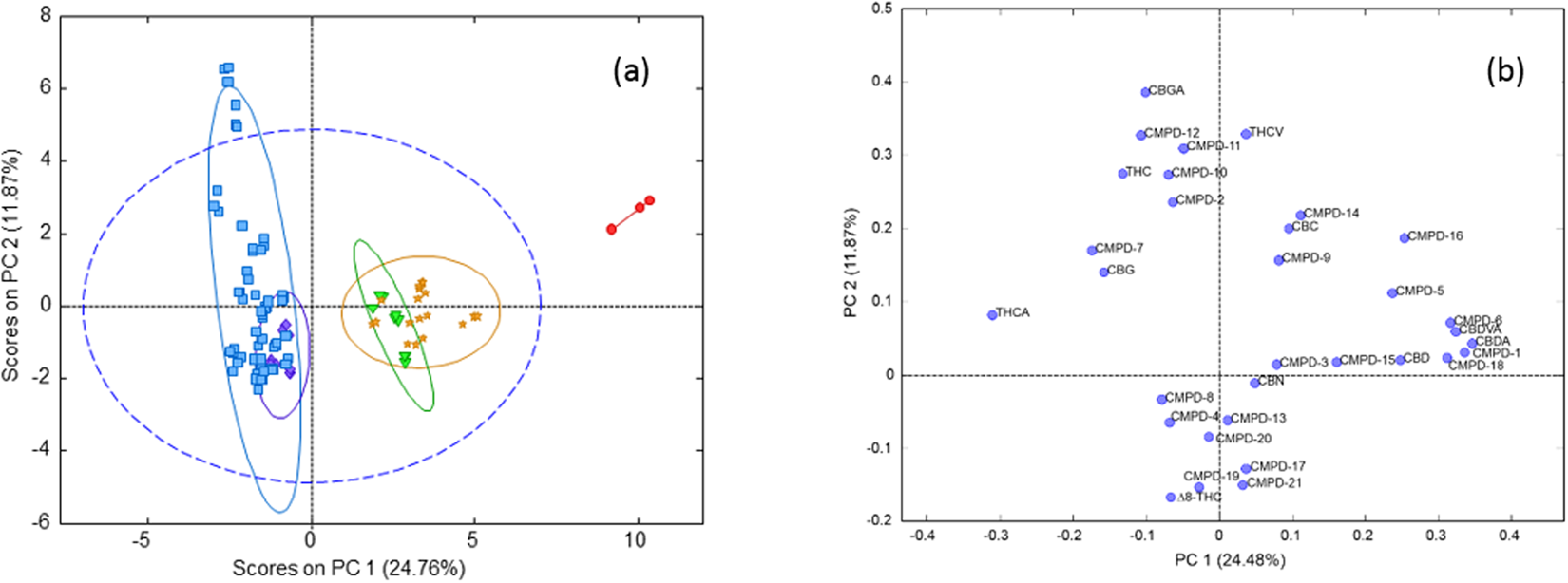
"Chemometric analysis of cannabinoids: Chemotaxonomy and domestication syndrome"
Cannabis is an interesting domesticated crop with a long history of cultivation and use. Strains have been selected through informal breeding programs with undisclosed parentage and criteria. The term “strain” refers to minor morphological differences and grower branding rather than distinct cultivated varieties. We hypothesized that strains sold by different licensed producers are chemotaxonomically indistinguishable and that the commercial practice of identifying strains by the ratio of total Δ9-tetrahydrocannabinol (THC) and cannabidiol (CBD) is insufficient to account for the reported human health outcomes. We used targeted metabolomics to analyze 11 known cannabinoids and an untargeted metabolomics approach to identify 21 unknown cannabinoids. Five clusters of chemotaxonomically indistinguishable strains were identified from the 33 commercial products. Only three of the clusters produce cannabidiolic acid (CBDA) in significant quantities, while the other two clusters redirect metabolic resources toward the tetrahydrocannabinolic acid (THCA) production pathways. (Full article...)
|
Featured article of the week: June 24–30:
"National and transnational security implications of asymmetric access to and use of biological data"
Biology and biotechnology have changed dramatically during the past 20 years, in part because of increases in computational capabilities and use of engineering principles to study biology. The advances in supercomputing, data storage capacity, and cloud platforms enable scientists throughout the world to generate, analyze, share, and store vast amounts of data, some of which are biological and much of which may be used to understand the human condition, agricultural systems, evolution, and environmental ecosystems. These advances and applications have enabled: (1) the emergence of data science, which involves the development of new algorithms to analyze and visualize data; and (2) the use of engineering approaches to manipulate or create new biological organisms that have specific functions, such as production of industrial chemical precursors and development of environmental bio-based sensors. Several biological sciences fields harness the capabilities of computer, data, and engineering sciences, including synthetic biology, precision medicine, precision agriculture, and systems biology. These advances and applications are not limited to one country. This capability has economic and physical consequences but is vulnerable to unauthorized intervention. (Full article...)
|
Featured article of the week: June 17–23:
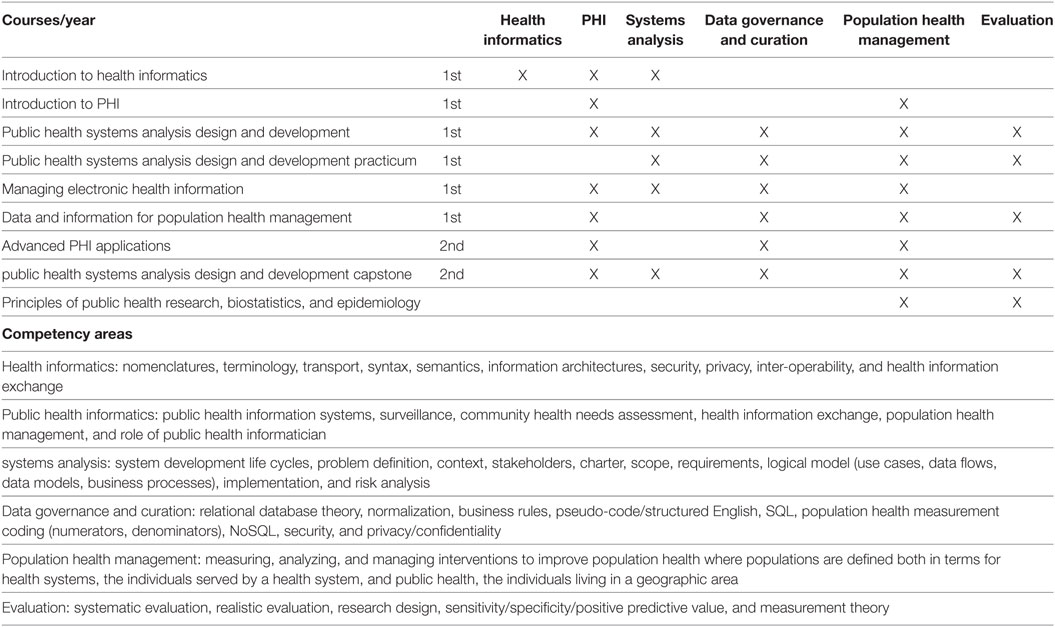
"Developing workforce capacity in public health informatics: Core competencies and curriculum design"
We describe a master’s level public health informatics (PHI) curriculum to support workforce development. Public health decision-making requires intensive information management to organize responses to health threats and develop effective health education and promotion. PHI competencies prepare the public health workforce to design and implement these information systems. The objective for a master's and certificate in PHI is to prepare public health informaticians with the competencies to work collaboratively with colleagues in public health and other health professions to design and develop information systems that support population health improvement. The PHI competencies are drawn from computer, information, and organizational sciences. A curriculum is proposed to deliver the competencies, and the results of a pilot PHI program are presented. Since the public health workforce needs to use information technology effectively to improve population health, it is essential for public health academic institutions to develop and implement PHI workforce training programs. (Full article...)
|
Featured article of the week: June 10–16:
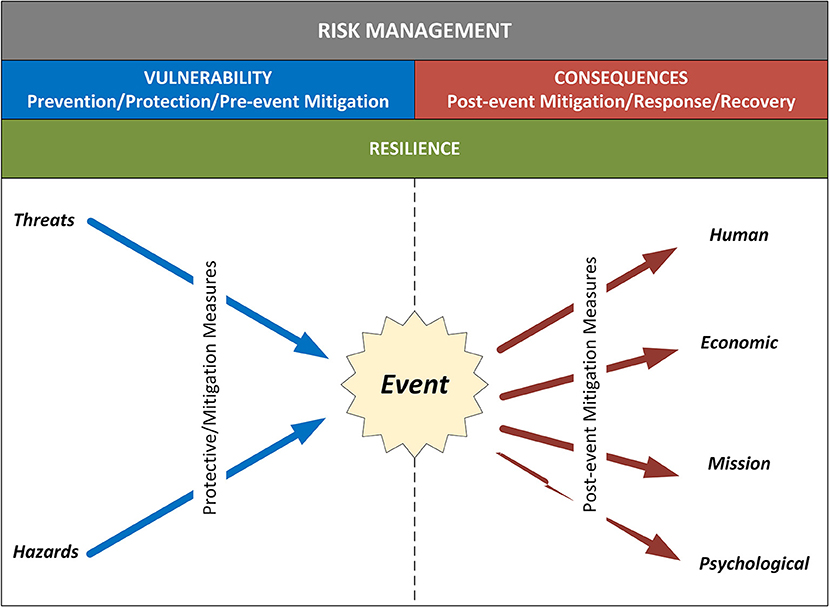
"Assessing cyberbiosecurity vulnerabilities and infrastructure resilience"
The convergence of advances in biotechnology with laboratory automation, access to data, and computational biology has democratized biotechnology and accelerated the development of new therapeutics. However, increased access to biotechnology in the digital age has also introduced additional security concerns and ultimately spawned the new discipline of cyberbiosecurity, which encompasses cybersecurity, cyber-physical security, and biosecurity considerations. With the emergence of this new discipline comes the need for a logical, repeatable, and shared approach for evaluating facility and system vulnerabilities to cyberbiosecurity threats. In this paper, we outline the foundation of an assessment framework for cyberbiosecurity, accounting for both security and resilience factors in the physical and cyber domains. This is a unique problem set, yet despite the complexity of the cyberbiosecurity field in terms of operations and governance, previous experience developing and implementing physical and cyber assessments applicable to a wide spectrum of critical infrastructure sectors provides a validated point of departure for a cyberbiosecurity assessment framework. (Full article...)
|
Featured article of the week: June 3–9:
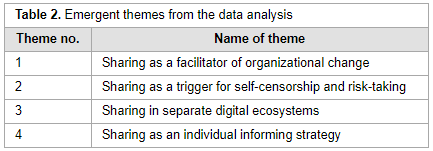
"What is the meaning of sharing: Informing, being informed or information overload?"
In recent years, several Norwegian public organizations have introduced enterprise social media platforms (ESMPs). The rationale for their implementation pertains to a goal of improving internal communications and work processes in organizational life. Such objectives can be attained on the condition that employees adopt the platform and embrace the practice of sharing. Although sharing work on ESMPs can bring benefits, making sense of the practice of sharing constitutes a challenge. In this regard, the paper performs an analysis on a case whereby an ESMP was introduced in a Norwegian public organization. The analytical focus is on the challenges and experiences of making sense of the practice of sharing. The research results show that users faced challenges in making sense of sharing. (Full article...)
|
Featured article of the week: May 27–June 2:
"Cyberbiosecurity: An emerging new discipline to help safeguard the bioeconomy"
Cyberbiosecurity is being proposed as a formal new enterprise which encompasses cybersecurity, cyber-physical security, and biosecurity as applied to biological and biomedical-based systems. In recent years, an array of important meetings and public discussions, commentaries, and publications have occurred that highlight numerous vulnerabilities. While necessary first steps, they do not provide a systematized structure for effectively promoting communication, education and training, elucidation, and prioritization for analysis, research, development, testing and evaluation, and implementation of scientific and technological standards of practice, policy, or regulatory or legal considerations for protecting the bioeconomy. Further, experts in biosecurity and cybersecurity are generally not aware of each other's domains, expertise, perspectives, priorities, or where mutually supported opportunities exist for which positive outcomes could result. (Full article...)
|
Featured article of the week: May 20–26:
"Cyberbiosecurity: A new perspective on protecting U.S. food and agricultural system"
Our national data and infrastructure security issues affecting the “bioeconomy” are evolving rapidly. Simultaneously, the conversation about cybersecurity of the U.S. food and agricultural system (cyber biosecurity) is incomplete and disjointed. The food and agricultural production sectors influence over 20% of the nation's economy ($6.7T) and 15% of U.S. employment (43.3M jobs). The food and agricultural sectors are immensely diverse, and they require advanced technologies and efficiencies that rely on computer technologies, big data, cloud-based data storage, and internet accessibility. There is a critical need to safeguard the cyber biosecurity of our bioeconomy, but currently protections are minimal and do not broadly exist across the food and agricultural system. Using the food safety management Hazard Analysis Critical Control Point (HACCP) system concept as an introductory point of reference, we identify important features in broad food and agricultural production and food systems: dairy, food animals, row crops, fruits and vegetables, and environmental resources (water). (Full article...)
|
Featured article of the week: May 13–19:
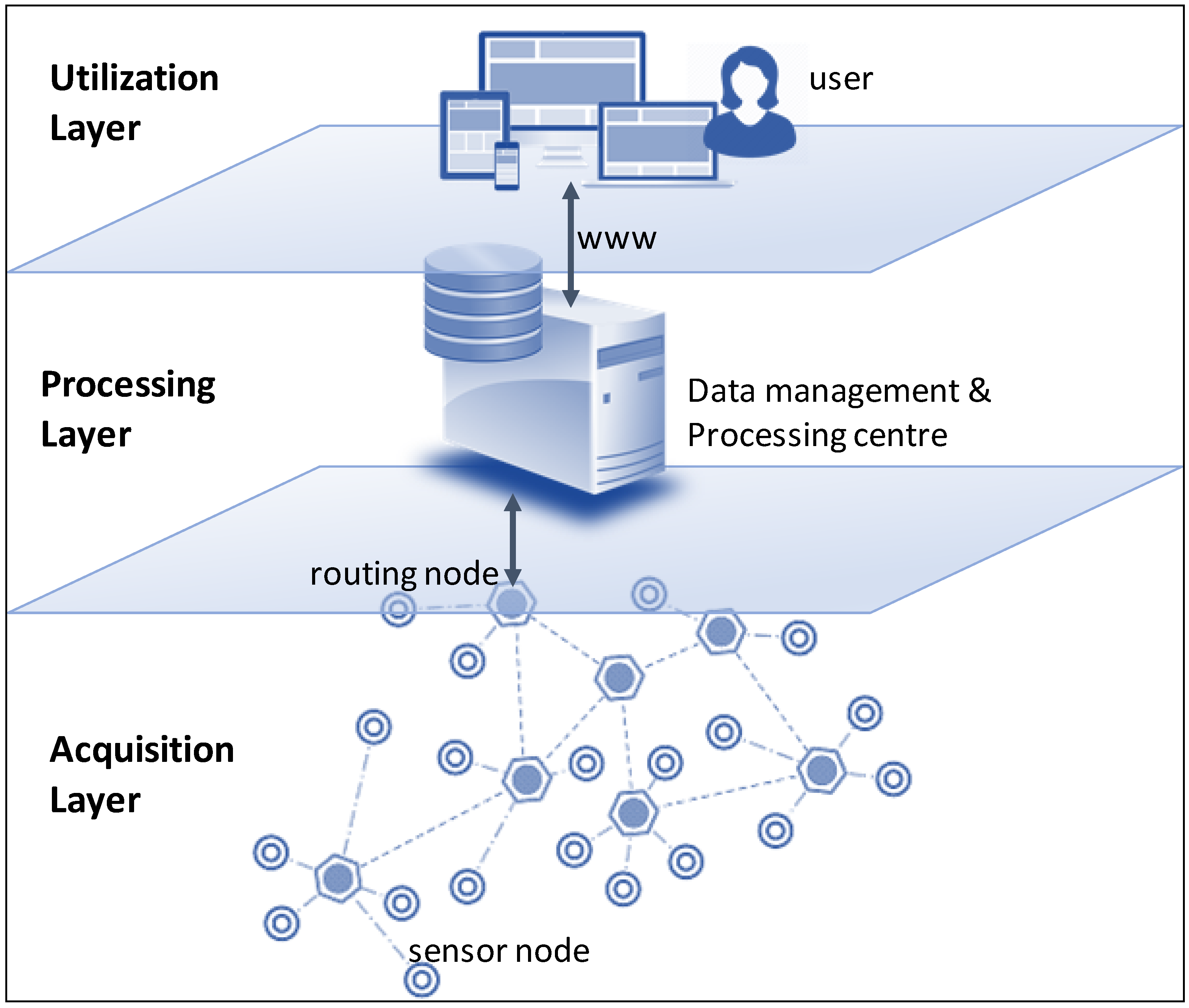
"DAQUA-MASS: An ISO 8000-61-based data quality management methodology for sensor data"
The internet of things (IoT) introduces several technical and managerial challenges when it comes to the use of data generated and exchanged by and between various smart, connected products (SCPs) that are part of an IoT system (i.e., physical, intelligent devices with sensors and actuators). Added to the volume and the heterogeneous exchange and consumption of data, it is paramount to assure that data quality levels are maintained in every step of the data chain/lifecycle. Otherwise, the system may fail to meet its expected function. While data quality (DQ) is a mature field, existing solutions are highly heterogeneous. Therefore, we propose that companies, developers, and vendors should align their data quality management mechanisms and artifacts with well-known best practices and standards, as for example, those provided by ISO 8000-61. This standard enables a process-approach to data quality management, overcoming the difficulties of isolated data quality activities. This paper introduces DAQUA-MASS, a methodology based on ISO 8000-61 for data quality management in sensor networks. (Full article...)
|
Featured article of the week: May 06–12:
"Security architecture and protocol for trust verifications regarding the integrity of files stored in cloud services"
Cloud computing is considered an interesting paradigm due to its scalability, availability, and virtually unlimited storage capacity. However, it is challenging to organize a cloud storage service (CSS) that is safe from the client point-of-view and to implement this CSS in public clouds since it is not advisable to blindly consider this configuration as fully trustworthy. Ideally, owners of large amounts of data should trust their data to be in the cloud for a long period of time, without the burden of keeping copies of the original data, nor of accessing the whole content for verification regarding data preservation. Due to these requirements, integrity, availability, privacy, and trust are still challenging issues for the adoption of cloud storage services, especially when losing or leaking information can bring significant damage, be it legal or business-related. With such concerns in mind, this paper proposes an architecture for periodically monitoring both the information stored in the cloud and the service provider behavior. (Full article...)
|
Featured article of the week: April 29–May 05:
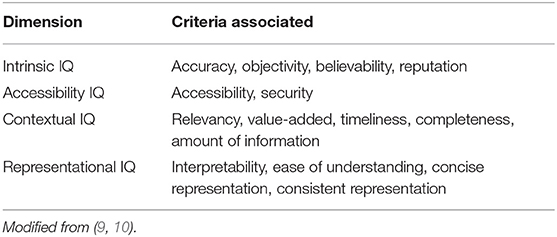
"What Is health information quality? Ethical dimension and perception by users"
The popularity of seeking health information online makes information quality (IQ) a public health issue. The present study aims at building a theoretical framework of health information quality (HIQ) that can be applied to websites and defines which IQ criteria are important for a website to be trustworthy and meet users' expectations. We have identified a list of HIQ criteria from existing tools and assessment criteria and elaborated them into a questionnaire that was promoted via social media and, mainly, the university. Responses (329) were used to rank the different criteria for their importance in trusting a website and to identify patterns of criteria using hierarchical cluster analysis. HIQ criteria were organized in five dimensions based on previous theoretical frameworks, as well as on how they cluster together in the questionnaire response. We could identify a top-ranking dimension (scientific completeness) that describes what the user is expecting to know from the websites (in particular: description of symptoms, treatments, side effects). (Full article...)
|
Featured article of the week: April 22–28:
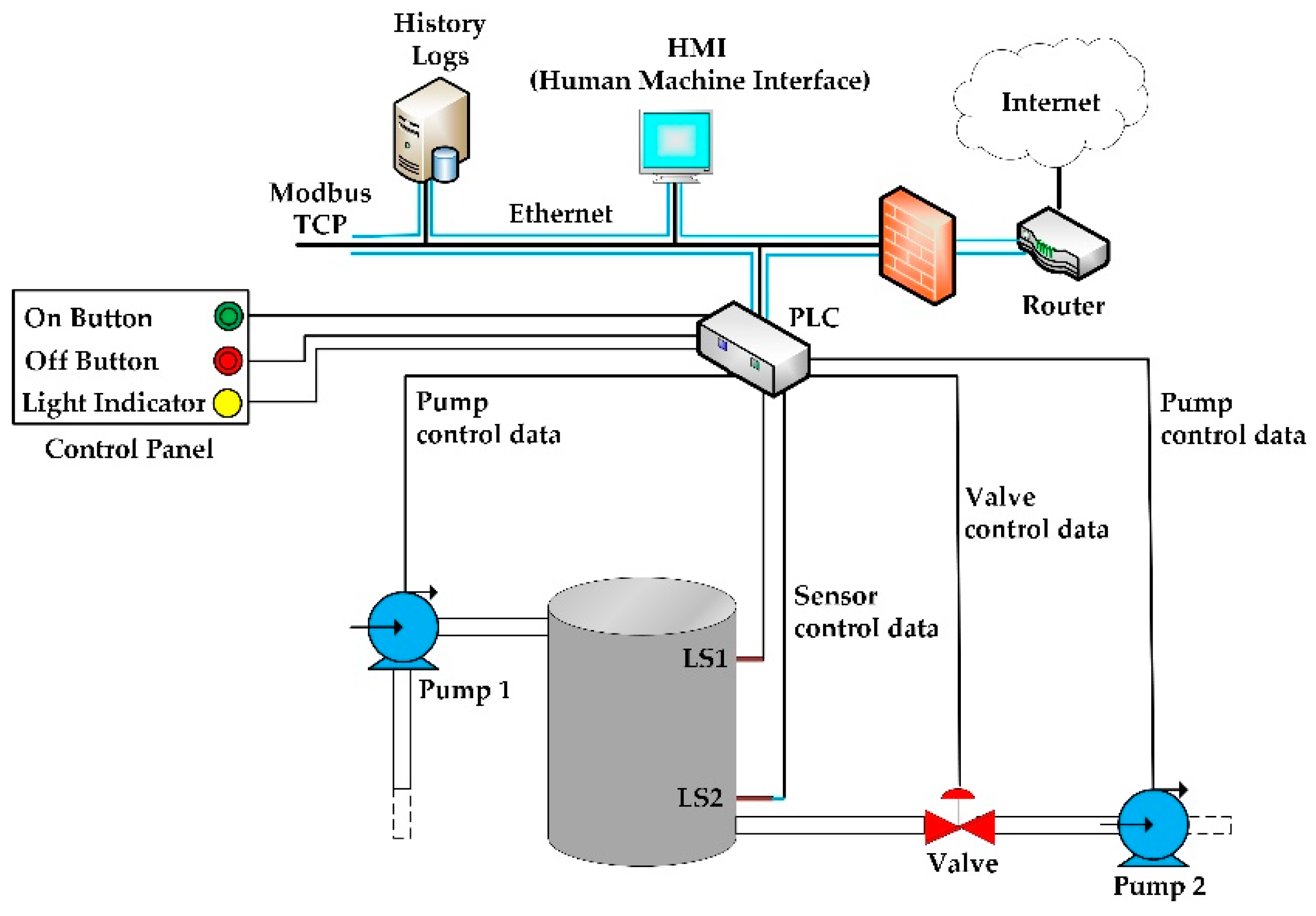
"SCADA system testbed for cybersecurity research using machine learning approach"
This paper presents the development of a supervisory control and data acquisition (SCADA) system testbed used for cybersecurity research. The testbed consists of a water storage tank’s control system, which is a stage in the process of water treatment and distribution. Sophisticated cyber-attacks were conducted against the testbed. During the attacks, the network traffic was captured, and features were extracted from the traffic to build a dataset for training and testing different machine learning algorithms. Five traditional machine learning algorithms were trained to detect the attacks: Random Forest, Decision Tree, Logistic Regression, Naïve Bayes, and KNN. Then, the trained machine learning models were built and deployed in the network, where new tests were made using online network traffic. The performance obtained during the training and testing of the machine learning models was compared to the performance obtained during the online deployment of these models in the network. The results show the efficiency of the machine learning models in detecting the attacks in real time. The testbed provides a good understanding of the effects and consequences of attacks on real SCADA environments. (Full article...)
|
Featured article of the week: April 15–21:
"Semantics for an integrative and immersive pipeline combining visualization and analysis of molecular data"
The advances made in recent years in the field of structural biology significantly increased the throughput and complexity of data that scientists have to deal with. Combining and analyzing such heterogeneous amounts of data became a crucial time consumer in the daily tasks of scientists. However, only few efforts have been made to offer scientists an alternative to the standard compartmentalized tools they use to explore their data and that involve a regular back and forth between them. We propose here an integrated pipeline especially designed for immersive environments, promoting direct interactions on semantically linked 2D and 3D heterogeneous data, displayed in a common working space. The creation of a semantic definition describing the content and the context of a molecular scene leads to the creation of an intelligent system where data are (1) combined through pre-existing or inferred links present in our hierarchical definition of the concepts, (2) enriched with suitable and adaptive analyses proposed to the user with respect to the current task and (3) interactively presented in a unique working environment to be explored. (Full article...)
|
Featured article of the week: April 8–14:
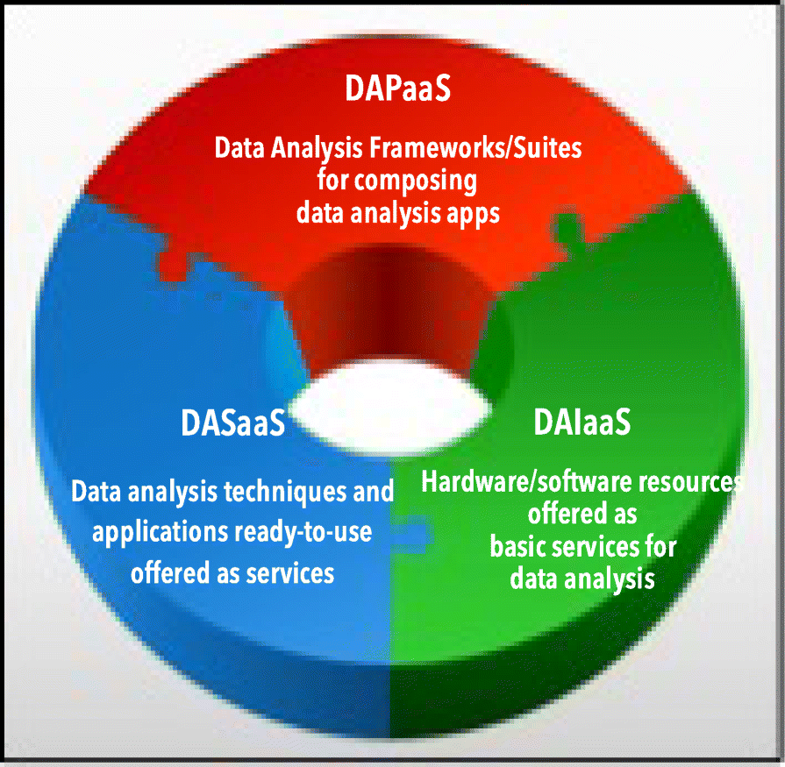
"A view of programming scalable data analysis: From clouds to exascale"
Scalability is a key feature for big data analysis and machine learning frameworks and for applications that need to analyze very large and real-time data available from data repositories, social media, sensor networks, smartphones, and the internet. Scalable big data analysis today can be achieved by parallel implementations that are able to exploit the computing and storage facilities of high-performance computing (HPC) systems and cloud computing systems, whereas in the near future exascale systems will be used to implement extreme-scale data analysis. Here is discussed how cloud computing currently supports the development of scalable data mining solutions and what the main challenges to be addressed and solved for implementing innovative data analysis applications on exascale systems currently are. (Full article...)
|
Featured article of the week: April 1–7:
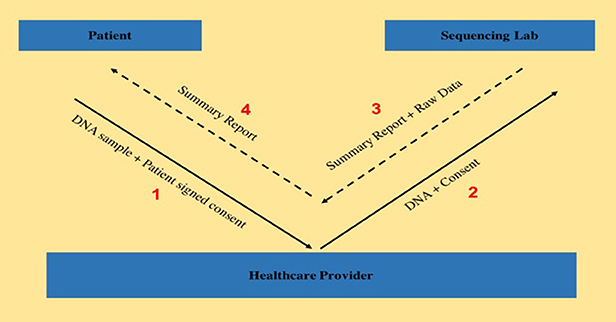
"Transferring exome sequencing data from clinical laboratories to healthcare providers: Lessons learned at a pediatric hospital"
The adoption rate of genome sequencing for clinical diagnostics has been steadily increasing, leading to the possibility of improvement in diagnostic yields. Although laboratories generate a summary clinical report, sharing raw genomic data with healthcare providers is equally important, both for secondary research studies as well as for a deeper analysis of the data itself, as seen by the efforts from organizations such as American College of Medical Genetics and Genomics, as well as Global Alliance for Genomics and Health. Here, we aim to describe the existing protocol of genomic data sharing between a certified clinical laboratory and a healthcare provider and highlight some of the lessons learned. This study tracked and subsequently evaluated the data transfer workflow for 19 patients, all of whom consented to be part of this research study and visited the genetics clinic at a tertiary pediatric hospital between April 2016 and December 2016. (Full article...)
|
Featured article of the week: March 25–31:
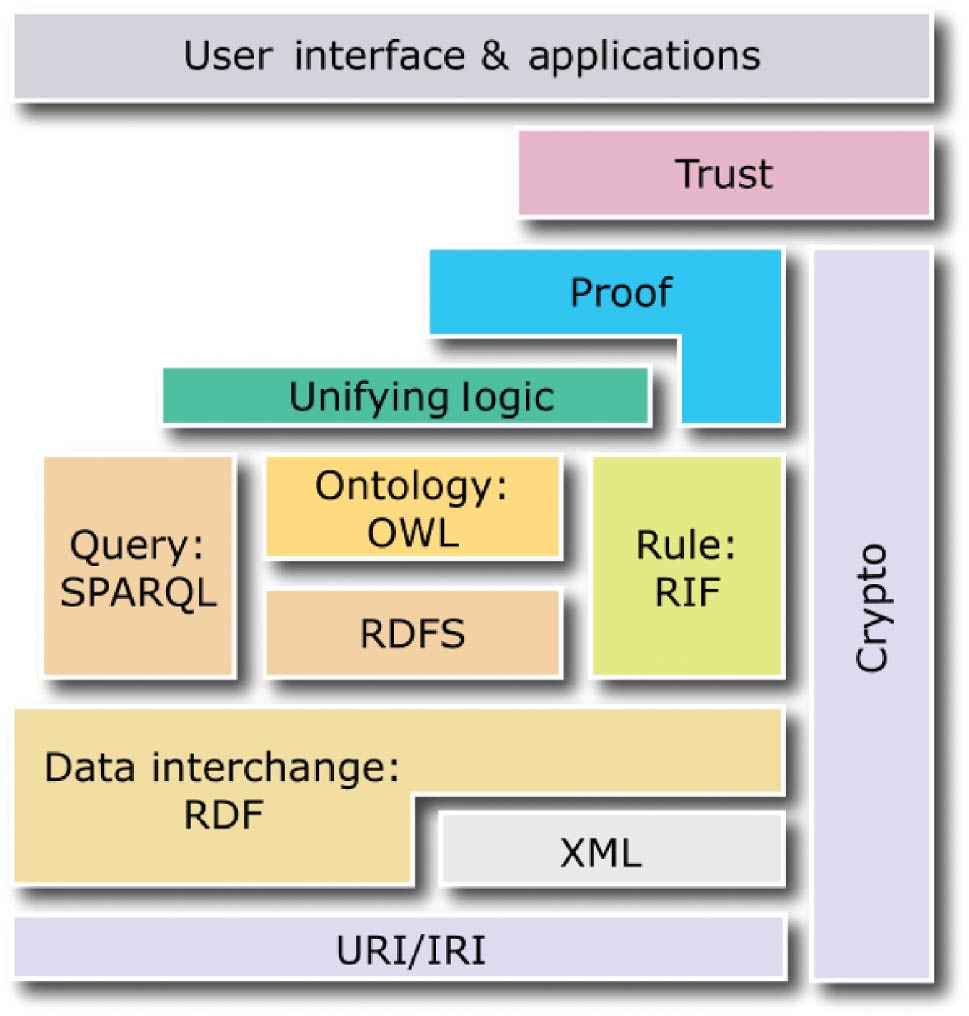
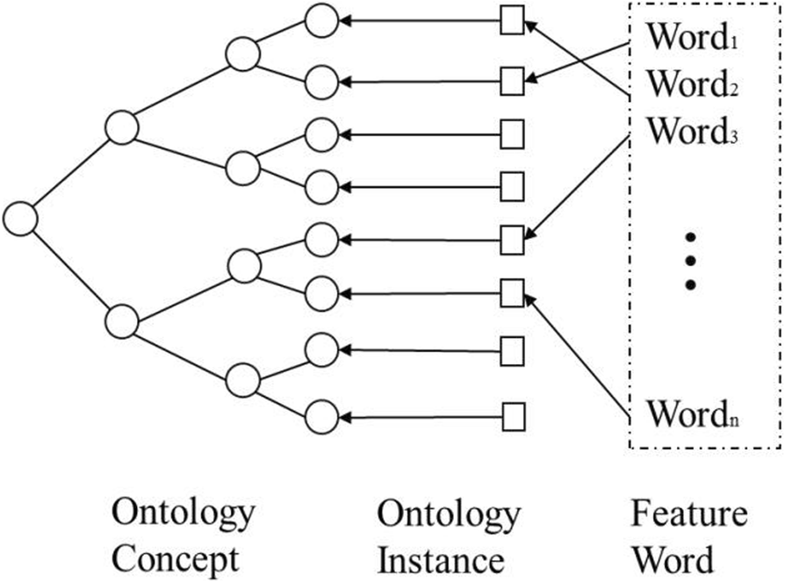
"Research on information retrieval model based on ontology"
An information retrieval system not only occupies an important position in the network information platform, but also plays an important role in information acquisition, query processing, and wireless sensor networks. It is a procedure to help researchers extract documents from data sets as document retrieval tools. The classic keyword-based information retrieval models neglect the semantic information which is not able to represent the user’s needs. Therefore, how to efficiently acquire personalized information that users need is of concern. The ontology-based systems lack an expert list to obtain accurate index term frequency. In this paper, a domain ontology model with document processing and document retrieval is proposed, and the feasibility and superiority of the domain ontology model are proved by the method of experiment. (Full article...)
|
Featured article of the week: March 18–24:
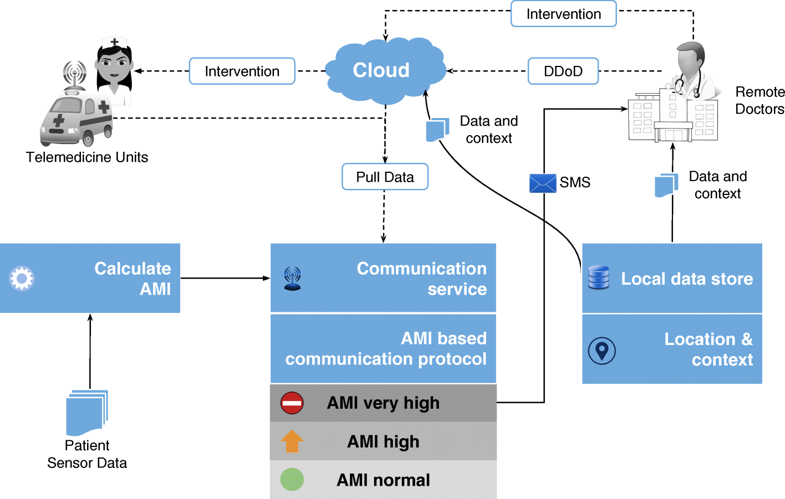
"Data to diagnosis in global health: A 3P approach"
With connected medical devices fast becoming ubiquitous in healthcare monitoring, there is a deluge of data coming from multiple body-attached sensors. Transforming this flood of data into effective and efficient diagnosis is a major challenge. To address this challenge, we present a "3P" approach: personalized patient monitoring, precision diagnostics, and preventive criticality alerts. In a collaborative work with doctors, we present the design, development, and testing of a healthcare data analytics and communication framework that we call RASPRO (Rapid Active Summarization for effective PROgnosis). The heart of RASPRO is "physician assist filters" (PAF) that 1. transform unwieldy multi-sensor time series data into summarized patient/disease-specific trends in steps of progressive precision as demanded by the doctor for a patient’s personalized condition, and 2. help in identifying and subsequently predictively alerting the onset of critical conditions. (Full article...)
|
Featured article of the week: March 11–17:
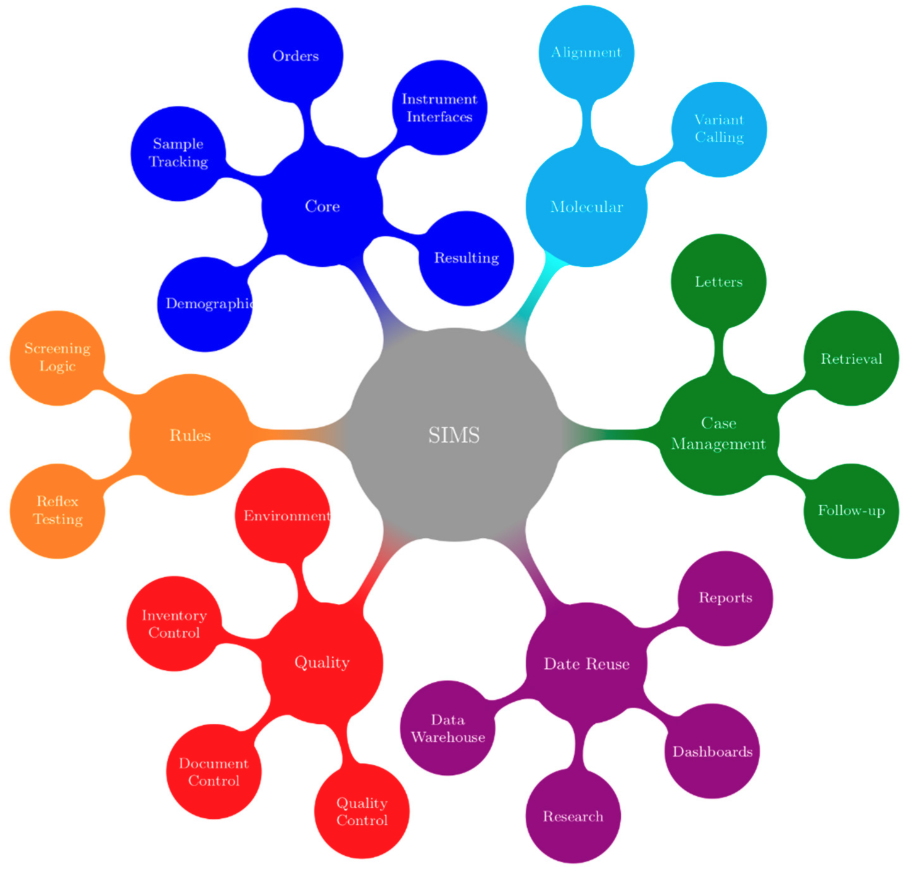
"Building a newborn screening information management system from theory to practice"
Information management systems are the central process management and communication hub for many newborn screening programs. In late 2014, Newborn Screening Ontario (NSO) undertook an end-to-end assessment of its information management needs, which resulted in a project to develop a flexible information systems (IS) ecosystem and related process changes. This enabled NSO to better manage its current and future workflow and communication needs. An idealized vision of a screening information management system (SIMS) was developed that was refined into enterprise and functional architectures. This was followed by the development of technical specifications, user requirements, and procurement. In undertaking a holistic full product lifecycle redesign approach, a number of change management challenges were faced by NSO across the entire program. Strong leadership support and full program engagement were key for overall project success. It is anticipated that improvements in program flexibility and the ability to innovate will outweigh the efforts and costs. (Full article...)
|
Featured article of the week: March 04–10:
"Adapting data management education to support clinical research projects in an academic medical center"
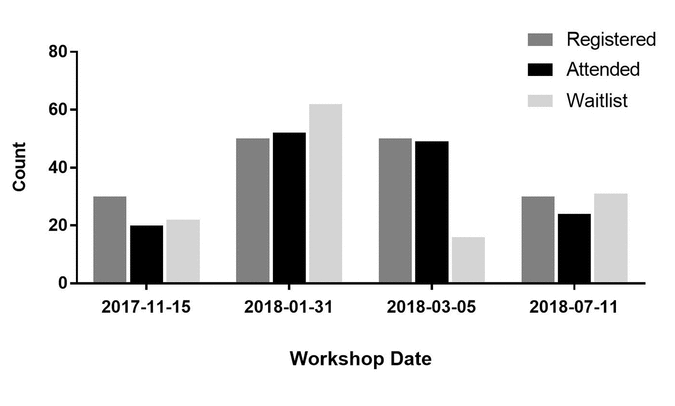
Librarians and researchers alike have long identified research data management (RDM) training as a need in biomedical research. Despite the wealth of libraries offering RDM education to their communities, clinical research is an area that has not been targeted. Clinical RDM (CRDM) is seen by its community as an essential part of the research process where established guidelines exist, yet educational initiatives in this area are unknown.
Leveraging my academic library’s experience supporting CRDM through informationist grants and REDCap training in our medical center, I developed a 1.5 hour CRDM workshop. This workshop was designed to use established CRDM guidelines in clinical research and address common questions asked by our community through the library’s existing data support program. The workshop was offered to the entire medical center four times between November 2017 and July 2018. This case study describes the development, implementation, and evaluation of this workshop. (Full article...)
|
Featured article of the week: February 25–March 03:
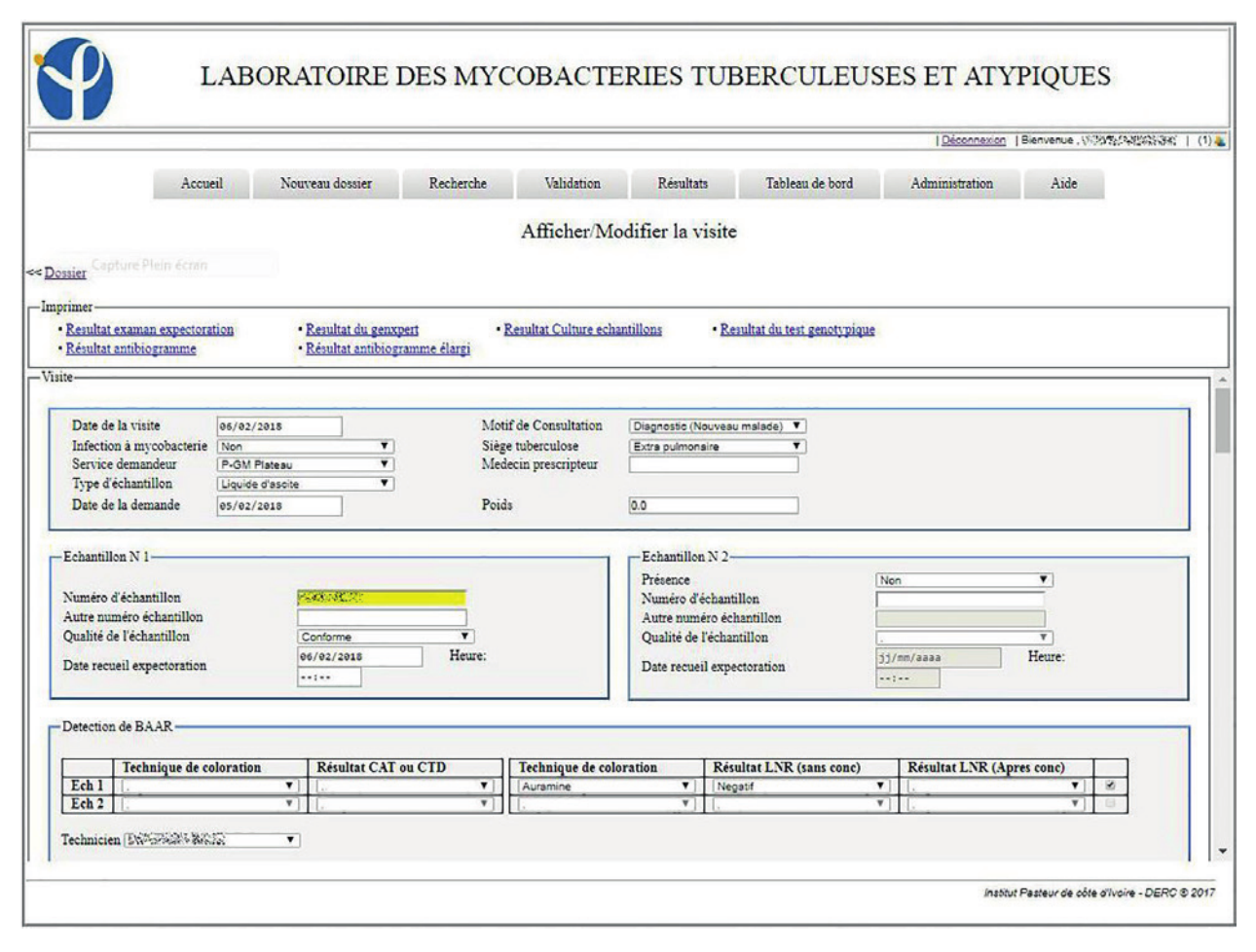
"Development of an electronic information system for the management of laboratory data of tuberculosis and atypical mycobacteria at the Pasteur Institute in Côte d’Ivoire"
Tuberculosis remains a public health problem despite all the efforts made to eradicate it. To strengthen the surveillance system for this condition, it is necessary to have a good data management system. Indeed, the use of electronic information systems in data management can improve the quality of data. The objective of this project was to set up a laboratory-specific electronic information system for mycobacteria and atypical tuberculosis.
The design of this laboratory information system required a general understanding of the workflow and the implementation processes in order to generate a realistic model. For the implementation of the system, Java technology was used to develop a web application compatible with the intranet of the company. (Full article...)
|
Featured article of the week: February 18–24:
"Codesign of the Population Health Information Management System to measure reach and practice change of childhood obesity programs"
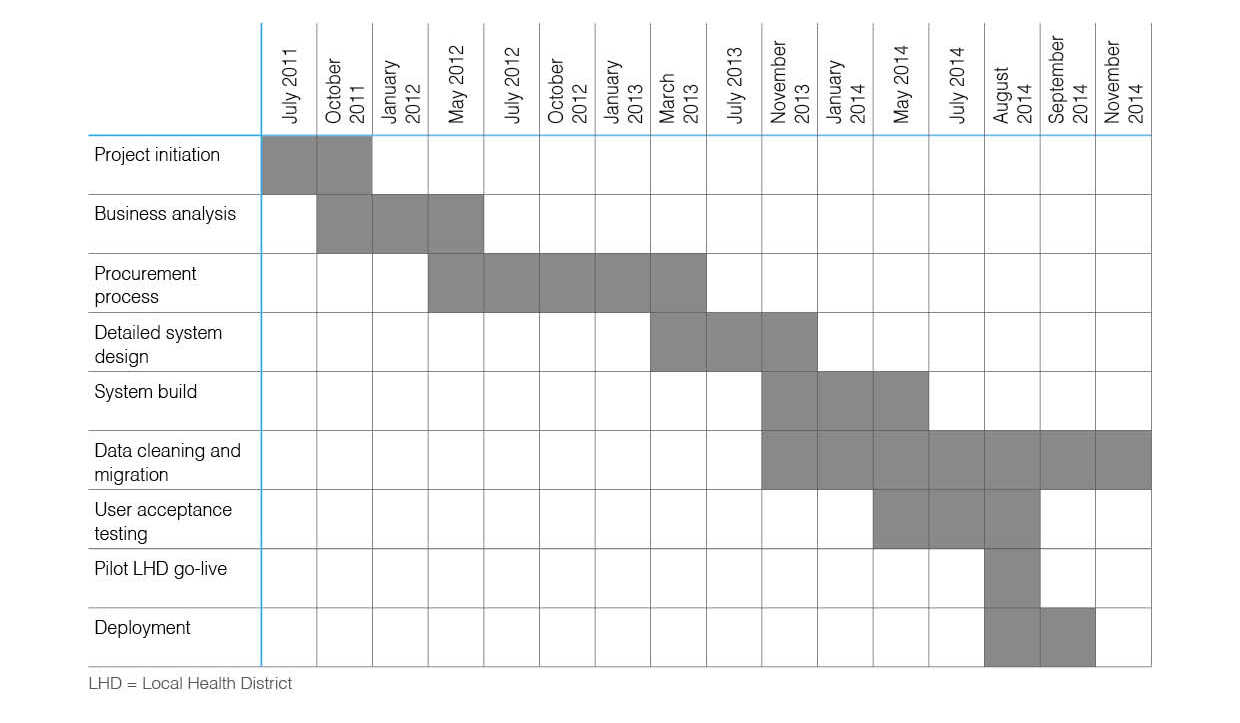
Childhood obesity prevalence is an issue of international public health concern, and governments have a significant role to play in its reduction. The Healthy Children Initiative (HCI) has been delivered in New South Wales (NSW), Australia, since 2011 to support implementation of childhood obesity prevention programs at scale. Consequently, a system to support local implementation and data collection, analysis, and reporting at local and state levels was necessary. The Population Health Information Management System (PHIMS) was developed to meet this need.
A collaborative and iterative process was applied to the design and development of the system. The process comprised identifying technical requirements, building system infrastructure, delivering training, deploying the system, and implementing quality measures. (Full article...)
|
Featured article of the week: February 11–17:
"Open data in scientific communication"
The development of information technology makes it possible to collect and analyze a growing number of data resources. The results of research, regardless of the discipline, constitute one of the main sources of data. Currently, research results are increasingly being published in the open access model. The open access concept has been accepted and recommended worldwide by many institutions financing and implementing research. Initially, the idea of openness concerned only the results of research and scientific publications; at present, more attention is paid to the problem of sharing scientific data, including raw data. Proceedings towards open data are intricate, as data specificity requires the development of an appropriate legal, technical and organizational model, followed by the implementation of data management policies at both the institutional and national levels. (Full article...)
|
Featured article of the week: February 4–10:
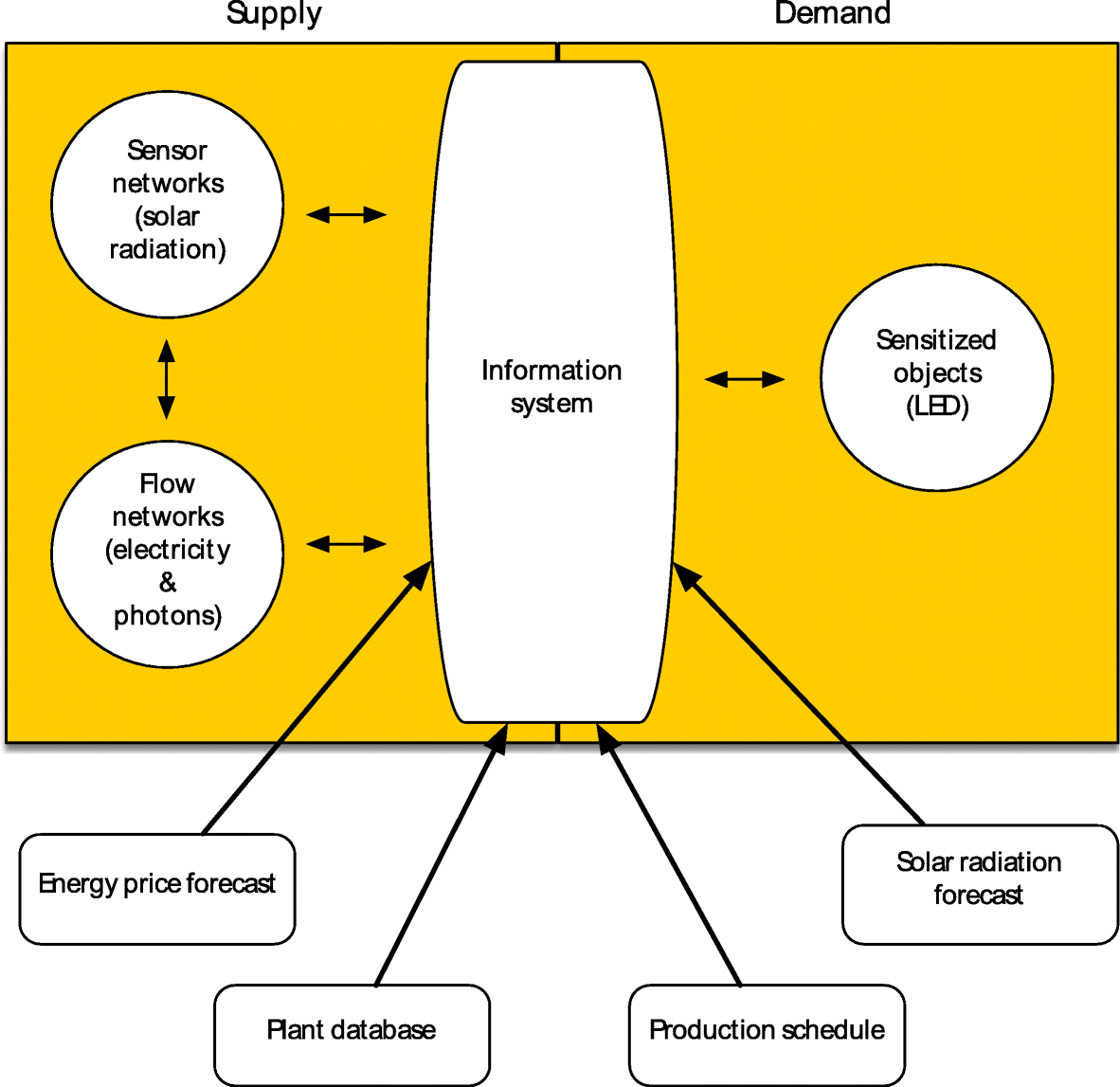
"Simulation of greenhouse energy use: An application of energy informatics"
Greenhouse agriculture is a highly efficient method of food production that can greatly benefit from supplemental electric lighting. The needed electricity associated with greenhouse lighting amounts to about 30% of its operating costs. As the light level of LED lighting can be easily controlled, it offers the potential to reduce energy costs by precisely matching the amount of supplemental light provided to current weather conditions and a crop’s light needs. Three simulations of LED lighting for growing lettuce in the Southeast U.S. using historical solar radiation data for the area were conducted. Lighting costs can be potentially reduced by approximately 60%. (Full article...)
|
Featured article of the week: January 28-February 3:
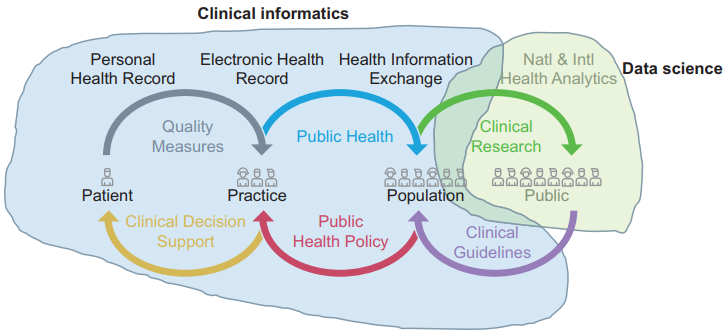
"Learning health systems need to bridge the "two cultures" of clinical informatics and data science"
United Kingdom (U.K.) health research policy and plans for population health management are predicated upon transformative knowledge discovery from operational "big data." Learning health systems require not only data but also feedback loops of knowledge into changed practice. This depends on knowledge management and application, which in turn depends upon effective system design and implementation. Biomedical informatics is the interdisciplinary field at the intersection of health science, social science, and information science and technology that spans this entire scope.
In the U.K., the separate worlds of health data science (bioinformatics, big data) and effective healthcare system design and implementation (clinical informatics, "digital health") have operated as "two cultures." Much National Health Service and social care data is of very poor quality. Substantial research funding is wasted on data cleansing or by producing very weak evidence. There is not yet a sufficiently powerful professional community or evidence base of best practice to influence the practitioner community or the digital health industry. (Full article...)
|
Featured article of the week: January 21-27:
"The problem with dates: Applying ISO 8601 to research data management"
Dates appear regularly in research data and metadata but are a problematic data type to normalize due to a variety of potential formats. This suggests an opportunity for data librarians to assist with formatting dates, yet there are frequent examples of data librarians using diverse strategies for this purpose. Instead, data librarians should adopt the international date standard ISO 8601. This standard provides needed consistency in date formatting, allows for inclusion of several types of date-time information, and can sort dates chronologically. As regular advocates for standardization in research data, data librarians must adopt ISO 8601 and push for its use as a data management best practice.(Full article...)
|
Featured article of the week: January 14–20:
"Health sciences libraries advancing collaborative clinical research data management in universities"
Medical libraries need to actively review their service models and explore partnerships with other campus entities to provide better-coordinated clinical research management services to faculty and researchers. TRAIL (Translational Research and Information Lab), a five-partner initiative at the University of Washington (UW), explores how best to leverage existing expertise and space to deliver clinical research data management (CRDM) services and emerging technology support to clinical researchers at UW and collaborating institutions in the Pacific Northwest. The initiative offers 14 services and a technology-enhanced innovation lab located in the Health Sciences Library (HSL) to support the University of Washington clinical and research enterprise. Sharing of staff and resources merges library and non-library workflows, better coordinating data and innovation services to clinical researchers. Librarians have adopted new roles in CRDM, such as providing user support and training for UW’s Research Electronic Data Capture (REDCap) instance. (Full article...)
|
Featured article of the week: January 7–13:
"Privacy preservation techniques in big data analytics: A survey"
Incredible amounts of data are being generated by various organizations like hospitals, banks, e-commerce, retail and supply chain, etc. by virtue of digital technology. Not only humans but also machines contribute to data streams in the form of closed circuit television (CCTV) streaming, web site logs, etc. Tons of data is generated every minute by social media and smart phones. The voluminous data generated from the various sources can be processed and analyzed to support decision making. However data analytics is prone to privacy violations. One of the applications of data analytics is recommendation systems, which are widely used by e-commerce sites like Amazon and Flipkart for suggesting products to customers based on their buying habits, leading to inference attacks. Although data analytics is useful in decision making, it will lead to serious privacy concerns. Hence privacy preserving data analytics became very important. This paper examines various privacy threats, privacy preservation techniques, and models with their limitations. The authors then propose a data lake-based modernistic privacy preservation technique to handle privacy preservation in unstructured data. (Full article...)
|
Featured article of the week: January 1–6:
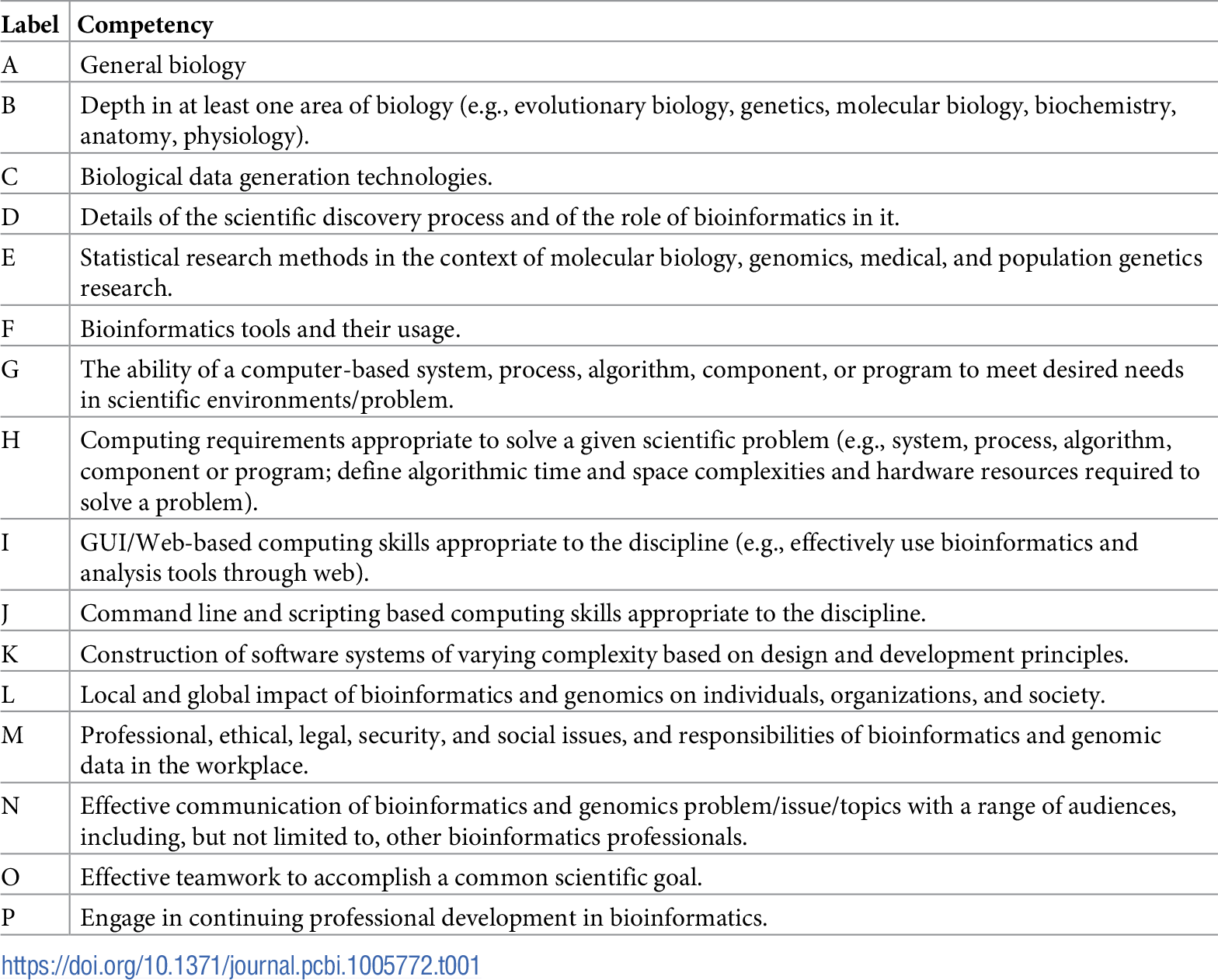
"The development and application of bioinformatics core competencies to improve bioinformatics training and education"
Bioinformatics is recognized as part of the essential knowledge base of numerous career paths in biomedical research and healthcare. However, there is little agreement in the field over what that knowledge entails or how best to provide it. These disagreements are compounded by the wide range of populations in need of bioinformatics training, with divergent prior backgrounds and intended application areas. The Curriculum Task Force of the International Society of Computational Biology (ISCB) Education Committee has sought to provide a framework for training needs and curricula in terms of a set of bioinformatics core competencies that cut across many user personas and training programs. The initial competencies developed based on surveys of employers and training programs have since been refined through a multiyear process of community engagement. This report describes the current status of the competencies and presents a series of use cases illustrating how they are being applied in diverse training contexts. (Full article...)
|
|