Journal:From command-line bioinformatics to bioGUI
| Full article title | From command-line bioinformatics to bioGUI |
|---|---|
| Journal | PeerJ |
| Author(s) | Joppich, Markus; Zimmer, Ralf |
| Author affiliation(s) | Ludwig-Maximilians-Universität München |
| Primary contact | Email: joppich at bio dot ifi dot lmu dot de |
| Editors | Gillespie, Joseph |
| Year published | 2019 |
| Volume and issue | 7 |
| Page(s) | e8111 |
| DOI | 10.7717/peerj.8111 |
| ISSN | 2167-8359 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://peerj.com/articles/8111/ |
| Download | https://peerj.com/articles/8111.pdf (PDF) |
|
|
This article should be considered a work in progress and incomplete. Consider this article incomplete until this notice is removed. |
Abstract
Bioinformatics is a highly interdisciplinary field providing informatics applications for scientists from many disciplines. Installing and starting applications on the command line (CL) is inconvenient and inefficient for many scientists. Nonetheless, most methods are implemented with a command-line interface only. Providing a graphical user interface (GUI) for bioinformatics applications is one step toward routinely making CL-only applications more readily available to scientists, yielding a positive step toward more effective interdisciplinary work. With our bioGUI framework, we address two main problems of using CL bioinformatics applications. First, many tools work on UNIX-based systems only, while many scientists use Microsoft Windows. Second, scientists refrain from using CL tools, which, despite their reservations, could well support them in their research. With bioGUI install modules and templates, installing and using CL tools is made possible for most scientists, even on Windows, due to bioGUI’s support for Windows Subsystem for Linux. In addition, bioGUI templates can easily be created, making the bioGUI framework highly rewarding for developers. From the bioGUI repository it is possible to download, install, and use bioinformatics tools with just a few clicks.
Introduction
Many advances in bioinformatics rely on sophisticated applications. Examples include Trinity[1] for de novo assembly in conjunction with Trimmomatic[2], or the HISAT2, StringTie, and Ballgown pipeline for transcript-level expression analysis.[3] These tools have in common that when locally installed, only a command-line interface (CLI) is provided, implying a burden for many conducting sequence analysis and alignments who are not computing-adept.[4] Jellyfish[5], Glimmer[6], and HMMER natively run only in UNIX-environments and require a sophisticated setup on Windows. In addition, the installation of command-line (CL) tools is a challenge for non-computer specialists, for example, due to package dependency resolution. This problem has been addressed by the AlgoRun package[7], providing a Docker-based repository of tools. Being a web-based service, it limits use to web-applicable data sizes, or local data must be made available to the Docker container in the cloud. While AlgoRun has the advantage of processing data anywhere, it relies on Docker. Docker may be run either on a local workstation or in the cloud. On a local workstation it can induce incompatibilities with existing software (using Hyper-V on Windows). A cloud-based service may conflict with data privacy guidelines[8], for example, with respect to a possible de-anonymization of patient samples.[9] Using Windows Subsystem for Linux (WSL) is often possible in such a scenario: it is provided as an app from the Microsoft Store.
A frequent argument for not providing a graphical user interface (GUI) is the overhead for developing it and the effort to make it truly “user centered.” Often GUIs are simply deemed unnecessary by application developers. However, one can be skeptical whether scientists who are not computing-adept can efficiently use CLIs in their research. In fact, bioinformatician Dr. István Albert[10] notes that “Bioinformatics, unfortunately, has quite the number of methods that represent the disconnect of the Ivory Tower.” Pavelin et al.[11] note that software is often developed without a focus on usability of interfaces (for end-users). While this does not imply that any GUI is helpful, we argue that without a GUI, the otherwise highly sophisticated CL applications are not very useful for some scientists. Besides, a GUI is often more convenient and helps to avoid using the wrong parameters, especially if an application is not yet routinely used in a lab. University of Western Ontario's David Roy Smith[12] also states that GUI-driven applications make daily work in biology or medical labs easier. Smith remarks that many end-users have a “penchant for point and click,” not being able to effectively use CL tools. Still, they should have the ability to access and analyse their own data. Many proprietary software solutions address this demand: they allow GUI-based data management, while also being extensible via plug-ins. Smith[13] also points out that one of the biggest advantages of such plugins is to combine the power of peer-reviewed algorithms with a user-friendly GUI. Thus, providing a GUI is an important step toward the applicability of methods by end-users.
Visne et al.[14] present a universal GUI for R aiming to close the gap between R developers and GUI-dependent users with limited R scripting skills. Additionally, web-based workflow systems, like Galaxy[15] or Yabi[16] provide a means to easily execute bioinformatics applications, but they tend to focus on more complex workflows. However, both Galaxy and Yabi are designed to be run and maintained by bioinformaticians for several users and are not meant to run on a single, individual basis, like in small labs. More recently Morais et al.[4] stated that the accessibility of bioinformatics applications is one of the main challenges of contemporary biology, and that one of the main problems for users is the struggle of using CLIs. While a GUI does not make an application user-friendly per se, it helps to make it more accessible by lowering the burden to use it.[14][4][17][18][19]
In recent Microsoft Windows operating systems the WSL feature can be activated. This feature provides a native, non-virtualized Ubuntu environment on Windows, allowing to run most applications that also run on Ubuntu. This solves the problems of running Unix-based tools on Windows. Remaining problems for scientists aiming to run bioinformatics applications thus might be the installation and usage of CL applications.
Here we present bioGUI, an open-source cross-platform framework for making CL applications more accessible via a GUI. It uses an XML-based domain-specific language (DSL) for template definition, which lowers the initial effort to create a GUI. bioGUI templates for CL applications can easily be scripted. Combined with install modules, the templates provide an efficient and convenient method to deploy bioinformatics applications on Microsoft Windows (via WSL), Mac OS, and Linux. bioGUI also addresses protocol/parameter management by saving completed templates, enabling easy reproducibility of data analyses (Fig. 1).
|
Methods
This section first summarizes existing GUI-based systems, then it covers the use-case study we performed and goes into detail of how bioGUI works.
Existing workflow systems
There are several workflow systems already available. Most prominent in bioinformatics are the Galaxy server and Yabi. In addition, workflow specification languages such as the Common Workflow Language (CWL) or Nextflow exist. These workflows do not directly compare to bioGUI because they (usually) require a server infrastructure and are not aimed to run on a local computer. However, they have in common that no CLI is needed to run bioinformatics applications.
With the R Gui Generator (RGG) a general GUI framework for R already exists. Recently, specialized GUI frameworks like SEED 2[19] or RNA CoMPASS[17] have been presented.
Galaxy and Yabi
The Galaxy server[15] is a well known workflow system in bioinformatics. While bioGUI does not aim to be a workflow system like Galaxy—for example, allowing data management—there are similarities. For instance, Galaxy also provides a web-based GUI for its workflows. However, all data to be processed by Galaxy must either be on the server itself or uploaded to a location that is reachable by the server. Galaxy can access cloud storage, but classified data may not be uploaded to such storage, as pointed out in the introduction. Additionally, Galaxy requires Unix knowledge to be installed and does not provide a binary for installation. Galaxy is not cross-platform compatible. (Microsoft Windows is supported through WSL but still requires Unix knowledge.) Galaxy users provide Docker containers for Galaxy, where a local storage can be mounted.
Another framework providing similar options is Yabi (Hunter et al., 2012). Yabi is only distributed using a Docker container.
Nextflow and DolphinNext
The combination of Nextflow and DolphinNext provides similar functionality to Galaxy or Yabi. While Nextflow is a DSL for describing general workflows (lacking a GUI definition), DolphinNext provides the web-based user interface (UI) which enables a convenient usage of Nextflow workflows. Nextflow requires a POSIX system architecture and may or may not run on Microsoft Windows using Cygwin. DolphinNext resembles a lot the Galaxy framework, which can make use of CWL workflows; however, it focuses on a deployment in a cluster environment. It is unknown whether or not both systems work on WSL.
Common Workflow Language
The CWL[20] is a new standard for workflow definition and defines a DSL. In this language, inputs, input-types, and the corresponding parameters are stored. Additionally, inputs can have a help text included.
Using the bio.tools ToolDog software[21], CWL workflows can be generated and exported for many bioinformatics applications. An advantage of using bio.tools is the automatic annotation and description of input and outputs. Unfortunately, for many packages no CWL workflows have been deposited.
SEED 2 and bioinformatics through windows
In contrast to the previously mentioned tools, SEED 2[19] and Bioinformatics through windows (BTW)[4] do not focus on running complex workflows in a cluster environment. Instead, these focus on specific tasks which can be run on regular laptop computers. SEED 2 focuses on amplicon high-throughput sequencing data analyses. On the other hand, BTW follows the same concept but focuses on the analysis of marker gene data and does not provide a GUI for this task. SEED 2 provides a GUI to perform the relevant analyses fast and conveniently, while BTW focuses on the usability of Unix CL tools on Windows.
RGG and AutoIt
RGG was developed as a general GUI framework for R applications.[14] It uses XML files to specify the input fields for the graphical representation. When the user has set all options, the GUI is translated into an R script for execution. The execution output can also be retrieved from the RGGRunner application. The RGG software is limited to R scripts, but the authors have expressed their hope that providing GUI for analytical pipelines could “help to bridge the gap between the R developer community and GUI-dependent users.”[14]
In contrast to RGG, AutoIt is a general automation framework which, similar to bioGUI, allows the definition of a GUI as well as a task that is executed according to this input. In contrast to AutoIt, bioGUI is cross-platform compatible, supports WSL, and provides install modules for bioinformatics applications.
Comparison of existing workflow systems to bioGUI
bioGUI is not a classical workflow system like Galaxy, CWL, or DolphinNext, when paired with Nextflow. bioGUI is not meant to run many tasks nor to run in a cluster environment. Moreover, bioGUI does not share the philosophy of having a compute cluster set up to run analyses in a repeated fashion. bioGUI is meant to enable the user to perform bioinformatics analysis at their work place. With bioGUI we aim to provide low-effort usage of bioinformatics applications, without the need to set up a complicated environment. This allows users to easily compare different methods on collected experimental data.
bioGUI finds its niche as a generalization of the concepts introduced by Větrovský, Baldrian, and Morais[19] and Morais et al.[4] SEED 2 provides a GUI such that a broad public has access to sophisticated and well-known bioinformatics CL applications in the context of amplicon analysis. Similar concepts, yet differently implemented, are provided by RNA CoMPASS[17] for pathogen-host transcriptome analysis or PipeCraft.[18] Here, custom web-based UIs let the user interact with their specialized pipelines. RGG[14] offers a general GUI framework for R applications only. bioGUI offers a similar framework, which is applicable to any Unix application. In both, RGG and bioGUI, users/developers specify the visual elements in an XML file. This XML file is then interpreted and translated into a GUI within an application (RGGRunner or bioGUI, respectively), which also shows the output of the script.
The bioGUI framework extends the concepts presented by RGG and SEED 2, for instance, to general applications, and improves accessibility to these applications by providing install modules.
Use case study
One of the main goals we had in mind when developing bioGUI was to create a powerful framework, which is easy-to-use for several types of users and that does not create significant overhead for the application developer. In order to study this, we introduce two classes of possible users. The first class represents a general user of the software who generally prefers a GUI for performing a research task, for example, data analysis after sequencing. The second class describes a software developer releasing an application of a new algorithm to solve the alignment of sequencing reads. This class thus depicts a typical developer.
From these two use cases (see also the Appendix section “Use cases”) we identify the requirements or goals for bioGUI:
- Installing new programs must be simple and should not require system administrators.
- Creating a GUI for a program must not take a lot of time.
- Templates must bring a basic GUI to run the programs, and output must not be interpreted.
- Templates must be saveable for later re-use and reference, and also be searchable.
- The system must be lightweight (runtime overhead, disk-space) to allow for running applications on laptops.
- Installing a program may require additional (protected) external files.
Finally, we developed a paper mock-up, with which we went through the anticipated workflow of the user. We identified several input components and features the bioGUI program has to include (Fig. A1, in the Appendix).
bioGUI approach
“The accessibility of bioinformatics applications is crucial and a challenge of contemporary biology.”[4] Particularly, the usage of CLIs poses a problem. Since most bioinformatics applications require the execution of commands on the CL for installation (such as for compilation, adding dependencies to the path variable, etc.), we estimate installation also poses a problem.
During the use-case study, and interviews with wet-lab scientists without a computational background (Q. Emslander, 2019, personal communication; L. Jimenez, 2019, personal communication), we found two main problems with bioinformatics applications for scientists, which we want to address with bioGUI: the installation of potentially useful applications and the application's usage. Both problems have in common that they are expected to be performed on the CL. A GUI for achieving the respective tasks in bioinformatics (and beyond) is missing.
In particular, the first task of installing bioinformatics applications on a user’s machine poses a few problems. Most bioinformatics applications are written for a Unix operating system, like Linux or Mac OS, while in general Microsoft Windows is the dominant operating system. In order to overcome this problem, bioGUI makes use of WSL on Windows. Even if the user’s OS is already Unix-like, using the CL to install software might prove cumbersome. Thus, in order to support all users, bioGUI uses a cross-platform approach. bioGUI is developed in C++ using the Qt framework.
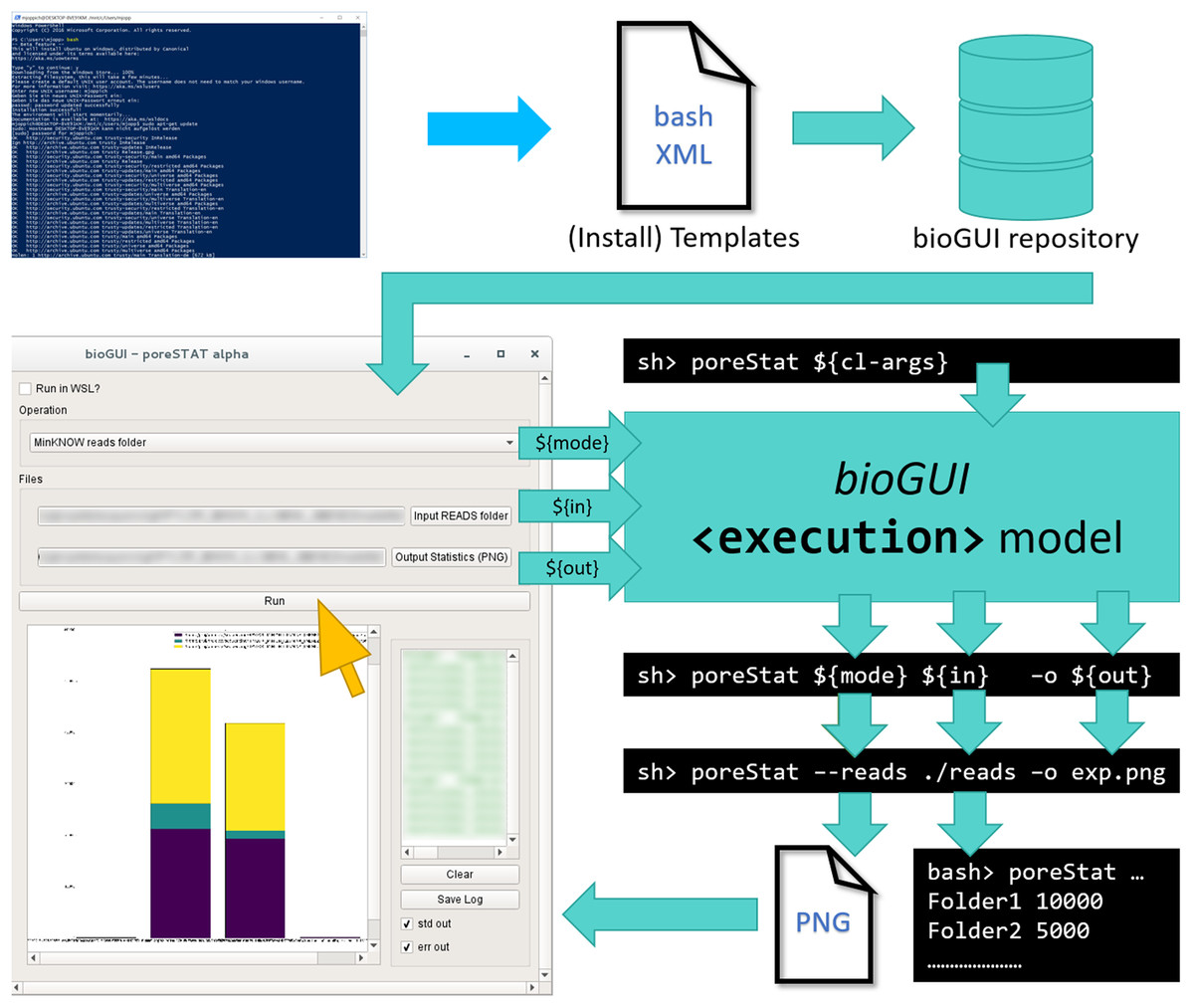
The general workflow for any program using bioGUI is shown in Fig. 1 (above). Given a CL application, the software developer (blue) writes the specific template in an XML-based DSL and can then make this template available, for example, in the bioGUI repository (cyan). Such templates can be automatically retrieved by bioGUI. Upon selection of a template by the user, bioGUI displays the input mask as defined in the template. When the user (yellow) has filled in all parameters, the parameters are collected by bioGUI and assembled into CL arguments, which are used to execute the original CL-only application. Upon completion, simple results (like text-output or images) can be shown in bioGUI directly or within an external application that is opened.
Install modules
Install modules are designed to install applications such that bioGUI can access them. Essentially, install modules are bash scripts which allow an automatic installation of applications into a predefined location. For this purpose, install modules receive several arguments from bioGUI when launched, for example, where to install the application to, the sudo password to fetch packages via a system’s package manager (e.g., aptitude, conda), whether the application should be made available to the user via the system’s PATH variable, etc. Install modules download and install applications and make them available to the user and bioGUI. However, some applications cannot be simply downloaded, but are distributed by installers. For this purpose, the install module template can be extended by further input fields. These must be specified by bioGUI elements, and their values are added to the end of the CL arguments of the install module. An install module can then execute the referenced installer.
Finally, an install module should contain the specification of its bioGUI template and could hardcode the path to the installed application. Other constant values, which can already be derived during the installation (e.g., absolute paths to dependencies), could also be defined in the template during this stage.
bioGUI templates
bioGUI templates are the actual end-user-interface to programs. A bioGUI template defines the look and functions of the UI. Thus it can define how the CL-application is called (with corresponding parameters).
Each bioGUI template consists of two parts (Fig. A2). The first part (the <window> model) defines the visual appearance of the GUI. The second part (the <execution> model) defines the processing logic of the template. Input values from the GUI components are collected and assembled (e.g., pre-/post-processing steps) to call CL applications. As part of this assembly, input values from the GUI may be transformed using multiple predefined nodes. Concatenations are possible using the <add> node, and constant values can be inserted using the <const> node. System environment properties, such as the operating system, the computer’s IP address, or specific directories can be collected using the <env> node. If the regular nodes are not sufficient, for example, because more complex string manipulations should be made (see use-case study), script nodes may also accept functions written in LUA or JavaScript.
In general, the execution part infers a network with inputs (e.g., GUI elements, other nodes within the execution part) and actions (if, add). For example, the execution network for an application with many sub-commands is exemplarily shown in Fig. A3 (see Appendix).
The time to template varies with the application as well as the number of options to be included. A simple template, like the one for MS-EmpiRe[22], can be created within 10 minutes. More comprehensive templates, like the one for HISAT2, usually take about 30 minutes. Time can be saved if only the most important command line options are shown in the GUI. This can be achieved by adding an “optional parameters” input field, where users can insert CL arguments themselves. This is, for instance, shown in the wtdbg2[23] and spades[24] templates. Adding the install part to a template usually can be done within 15 to 30 minutes, depending on how detailed the build process is documented. The creation of an install module thus takes approximately one hour.
bioGUI integration with CWL and argparse
The CWL[20] only describes the CL workflow and neither provides a GUI nor a means to install the desired tool. Due to this more general specification, CWL fits most problems, but specific annotations of inputs, explanations, or the embedding of images is not supported in CWL.
While developers can always create templates manually, bioGUI supports developers by offering a template generator from CWL templates or python3 argparse CL parsers. Since there are already many CWL templates available for bioinformatics CL applications, CWL files can be used as a base to automatically generate bioGUI templates from. Using the bioGUI template generator for argparse, it is also possible to automatically generate templates from CWL files (making use of the cwl2argparse program provided by CWL). Our generator takes as input the argparse parser or CWL file and creates input elements for all elements. In case the type of an input is unclear or not supported, the generator falls back to a regular text-input element.
Results
bioGUI templates
Currently more than 25 install modules exist for bioGUI. These represent basically three groups of bioinformatics tasks: next-generation sequencing data analysis and transcriptomics, long read sequencing analysis and assembly, and a more general sequence analysis. In general, these install modules will install the respective application on the local machine. The Circlator[25] template allows to pull and use the corresponding Docker image. The available tools, as well as their respective categorization, are listed in Table 1.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Supplemental information
- DOI 10.7717/peerj.8111/supp-1 - Survey questions on command-line tools and bioGUI: This is the original survey used to assess problems with current bioinformatics applications. (PDF)
- DOI 10.7717/peerj.8111/supp-2 - Answers on the survey on command-line tools and bioGUI: Each column represents a single participant. Questions are in rows. (XLXS)
Acknowledgements
We thank Luisa F. Jimenez-Soto and Gergely Csaba for their valuable input as well as for reviewing the manuscript. We thank the participants in our survey for their time. We thank the reviewers for their constructive feedback.
Authors’ contributions
Markus Joppich conceived and designed the experiments, performed the experiments, analyzed the data, prepared figures and/or tables, authored or reviewed drafts of the paper, and approved the final draft. Ralf Zimmer conceived and designed the experiments, analyzed the data, prepared figures and/or tables, authored or reviewed drafts of the paper, and approved the final draft.
Funding
This work was supported by the Deutsche Forschungsgemeinschaft (Collaborative Research Centre SFB 1123-2/Z2). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Data availability
The bioGUI documentation is available here. In order to set up Windows Subsystem for Linux (required for using bioGUI on Windows), follow the steps documented here. bioGUI is open-source software. Releases and code are available on the GitHub project page. Additional software (cwl2biogui) is available here.
Competing interests
The authors declare that they have no competing interests.
References
- ↑ Grabherr, M.G.; Haas, B.J.; Yassour, M. et al. (2011). "Full-length transcriptome assembly from RNA-Seq data without a reference genome". Nature Biotechnology 29 (7): 644–52. doi:10.1038/nbt.1883. PMC PMC3571712. PMID 21572440. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3571712.
- ↑ Bolger, A.M.; Lohse, M.; Usadel, B. (2014). "Trimmomatic: A flexible trimmer for Illumina sequence data". Bioinformatics 30 (15): 2114-20. doi:10.1093/bioinformatics/btu170. PMC PMC4103590. PMID 24695404. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4103590.
- ↑ 3.0 3.1 Pertea, M.; Kim, D.; Pertea, G.M. et al. (2016). Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. 11. pp. 1650–67. doi:10.1038/nprot.2016.095. PMC PMC5032908. PMID 27560171. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5032908.
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 Morais, D.; Roesch, L.F.W.; Redmile-Gordon, M. et al. (2018). BTW-Bioinformatics Through Windows: An easy-to-install package to analyze marker gene data. 6. pp. e5299. doi:10.7717/peerj.5299. PMC PMC6074753. PMID 30083449. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6074753.
- ↑ Marçais, G.; Kingsford, C. (2011). "A fast, lock-free approach for efficient parallel counting of occurrences of k-mers". Bioinformatics 27 (6): 764–70. doi:10.1093/bioinformatics/btr011. PMC PMC3051319. PMID 21217122. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3051319.
- ↑ 6.0 6.1 Delcher, A.L.; Bratke, K.A.; Powers, E.C. et al. (2007). "Identifying bacterial genes and endosymbiont DNA with Glimmer". Bioinformatics 23 (6): 673–9. doi:10.1093/bioinformatics/btm009. PMC PMC2387122. PMID 17237039. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2387122.
- ↑ Hosny, A.; Vera-Licona, P.; Laubenbacher, R. et al. (2016). "AlgoRun: A Docker-based packaging system for platform-agnostic implemented algorithms". Bioinformatics 32 (15): 2396–8. doi:10.1093/bioinformatics/btw120. PMC PMC6280798. PMID 27153722. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6280798.
- ↑ Schadt, E.E. (2012). "The changing privacy landscape in the era of big data". Molecular Systems Biology 8: 612. doi:10.1038/msb.2012.47. PMC PMC3472686. PMID 22968446. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3472686.
- ↑ Gymrek, M.; McGuire, A.L.; Golan, D. et al. (2013). "Identifying personal genomes by surname inference". Science 339 (6117): 321–4. doi:10.1126/science.1229566. PMID 23329047.
- ↑ Albert, I. (2016). "The Biostar Handbook". https://biostar.myshopify.com/.
- ↑ Pavelin, K.; Cham, J.A.; de Matos, P. et al. (2012). "Bioinformatics meets user-centred design: a perspective". PLoS Computational Biology 8 (7): e1002554. doi:10.1371/journal.pcbi.1002554. PMC PMC3395592. PMID 22807660. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3395592.
- ↑ Smith, D.R. (2013). "The battle for user-friendly bioinformatics". Frontiers in Genetics 4: 187. doi:10.3389/fgene.2013.00187. PMC PMC3778374. PMID 24065986. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3778374.
- ↑ Smith, D.R. (2015). Buying in to bioinformatics: An introduction to commercial sequence analysis software. 16. p. 700-9. doi:10.1093/bib/bbu030. PMC PMC4501248. PMID 25183247. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4501248.
- ↑ 14.0 14.1 14.2 14.3 14.4 Visne, I.; Dilaveroglu, E.; Vierlinger, K. et al. (2009). RGG: A general GUI Framework for R scripts. 10. p. 74. doi:10.1186/1471-2105-10-74. PMC PMC2653488. PMID 19254356. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2653488.
- ↑ 15.0 15.1 Afgan, E.; Baker, D.; van den Beek, M. et al. (2016). "The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update". Nucleic Acids Research 44 (W1): W3–W10. doi:10.1093/nar/gkw343. PMC PMC4987906. PMID 27137889. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4987906.
- ↑ Hunter, A.A.; Macgregor, A.B.; Szabo, T.O. et al. (2012). "Yabi: An online research environment for grid, high performance and cloud computing". Source Code for Biology and Medicine 7 (1): 1. doi:10.1186/1751-0473-7-1. PMC PMC3298538. PMID 22333270. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3298538.
- ↑ 17.0 17.1 17.2 Xu, G.; Strong, M.J.; Lacey, M.R. et al. (2014). "RNA CoMPASS: A dual approach for pathogen and host transcriptome analysis of RNA-seq datasets". PLoS One 9 (2): e89445. doi:10.1371/journal.pone.0089445. PMC PMC3934900. PMID 24586784. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3934900.
- ↑ 18.0 18.1 Anslan, S.; Bahram, M.; Hiirsalu, I. et al. (2017). "PipeCraft: Flexible open-source toolkit for bioinformatics analysis of custom high-throughput amplicon sequencing data". Molecular Ecology Resources 17 (6): e234-e240. doi:10.1111/1755-0998.12692. PMID 28544559.
- ↑ 19.0 19.1 19.2 19.3 Vetrovský, T.; Baldrian, P.; Morais, D. (2018). "SEED 2: A user-friendly platform for amplicon high-throughput sequencing data analyses". Bioinformatics 34 (13): 2292-2294. doi:10.1093/bioinformatics/bty071. PMC PMC6022770. PMID 29452334. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6022770.
- ↑ 20.0 20.1 Amstutz, P.; Andeer, R.; Chapman, B. et al. (15 March 2016). "Common Workflow Language, Draft 3". FigShare. https://figshare.com/articles/Common_Workflow_Language_draft_3/3115156/1.
- ↑ Hillion KH1, Kuzmin I2, Khodak A. et al. (2017). "Using bio.tools to generate and annotate workbench tool descriptions". F1000Research 6: ELIXIR-2074. doi:10.12688/f1000research.12974.1. PMC PMC5747335. PMID 29333231. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5747335.
- ↑ Ammar, C.; Gruber, M.; Csaba, G. et al. (2019). "MS-EmpiRe Utilizes Peptide-level Noise Distributions for Ultra-sensitive Detection of Differentially Expressed Proteins". Molecular and Cellular Proteomics 18 (9): 1880–92. doi:10.1074/mcp.RA119.001509. PMC PMC6731086. PMID 31235637. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6731086.
- ↑ Ruan, J.; Li, H. (2019). "Fast and accurate long-read assembly with wtdbg2". Nature Methods. doi:10.1038/s41592-019-0669-3. PMID 31819265.
- ↑ Bankevich, A.; Nurk, S.; Antipov, D. et al. (2012). "SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing". Journal of Computational Biology. doi:10.1089/cmb.2012.0021. PMC PMC3342519. PMID 22506599. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3342519.
- ↑ Hunt, M.; Silva, N.D.; Otto, T.D. et al. (2015). "Circlator: Automated circularization of genome assemblies using long sequencing reads". Genome Biology 16: 294. doi:10.1186/s13059-015-0849-0. PMC PMC4699355. PMID 26714481. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4699355.
- ↑ Langmead, B.; Trapnell, C.; Pop, M. et al. (2009). "Ultrafast and memory-efficient alignment of short DNA sequences to the human genome". Genome Biology 10 (3): R25. doi:10.1186/gb-2009-10-3-r25. PMC PMC2690996. PMID 19261174. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2690996.
- ↑ Langmead, B.; Salzberg, S.L. (2012). "Fast gapped-read alignment with Bowtie 2". Mature Methods 9 (4): 357–9. doi:10.1038/nmeth.1923. PMC PMC3322381. PMID 22388286. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3322381.
- ↑ Li, H.; Durbin, R. (2009). "Fast and accurate short read alignment with Burrows-Wheeler transform". Bioinformatics 25 (14): 1754–60. doi:10.1093/bioinformatics/btp324. PMC PMC2705234. PMID 19451168. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2705234.
- ↑ Koren, S.; Walenz, B.P.; Berlin, K. et al. (2017). "Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation". Genome Research 27 (5): 722–36. doi:10.1101/gr.215087.116. PMC PMC5411767. PMID 28298431. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5411767.
- ↑ Liao, Y.; Smyth, G.K.; Shi, W. (2014). "featureCounts: an efficient general purpose program for assigning sequence reads to genomic features". Bioinformatics 30 (7): 923–30. doi:10.1093/bioinformatics/btt656. PMID 24227677.
- ↑ Sović, I.; Šikić, M.; Wilm, A. et al. (2016). "Fast and sensitive mapping of nanopore sequencing reads with GraphMap". Nature Communications 7: 11307. doi:10.1038/ncomms11307. PMC PMC4835549. PMID 27079541. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4835549.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation, spelling, and grammar. We also added PMCID and DOI when they were missing from the original reference. The original article lists references alphabetically, but this version—by design—lists them in order of appearance.