Journal:Laboratory information system requirements to manage the COVID-19 pandemic: A report from the Belgian national reference testing center
| Full article title | Laboratory information system requirements to manage the COVID-19 pandemic: A report from the Belgian national reference testing center |
|---|---|
| Journal | Journal of the American Medical Informatics Association |
| Author(s) |
Weemaes, Matthias; Martens, Steven; Cuypers, Lize; Van Elslande, Jan; Hoet, Katrien; Welkenhuysen, Joris; Goossens, Ria; Wouters, Stijn; Houben, Els; Jeuris, Kirsten; Laenen, Lies; Bruyninckx, Katrien; Beuselinck, Kurt; André, Emmanuel; Depypere, Melissa; Desmet, Stefanie; Lagrou, Katrien; Van Ranst, Marc; Verdonck, Ann K.L.C.; Goveia, Jermaine |
| Author affiliation(s) | University Hospitals Leuven, Katholieke Universiteit Leuven |
| Primary contact | Email: jermaine dot goveia at uzleuven dot be |
| Year published | 2020 |
| Volume and issue | Ahead of print |
| Article # | ocaa081 |
| DOI | 10.1093/jamia/ocaa081 |
| ISSN | 1527-974X |
| Distribution license | Creative Commons Attribution Non-Commercial 4.0 International |
| Website | https://academic.oup.com/jamia/advance-article/doi/10.1093/jamia/ocaa081/5827002 |
| Download | https://academic.oup.com/jamia/advance-article-pdf/doi/10.1093/jamia/ocaa081/33421026/ocaa081.pdf (PDF) |
|
|
This article should be considered a work in progress and incomplete. Consider this article incomplete until this notice is removed. |
Abstract
Objective: The study sought to describe the development, implementation, and requirements of laboratory information system (LIS) functionality to manage test ordering, registration, specimen flow, and result reporting during the coronavirus disease 2019 (COVID-19) pandemic.
Materials and methods: Our large (more than 12,000,000 tests/year) academic hospital laboratory is the Belgian National Reference Center for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) testing. We have performed a moving total of more than 25,000 SARS-CoV-2 polymerase chain reaction tests in parallel to standard routine testing since the start of the outbreak. A LIS implementation team dedicated to developing tools to remove workflow bottlenecks—primarily situated in the pre- and post-analytical phases—was established early in the crisis.
Results: We outline the design, implementation, and requirements of LIS functionality related to managing increased test demand during the COVID-19 crisis, including tools for test ordering, standardized order sets integrated into a computerized physician order entry module, notifications on shipping requirements, automated triaging based on digital metadata forms, and the establishment of databases with contact details of other laboratories and primary care physicians to enable automated reporting. We also describe our approach to data mining and reporting of actionable daily summary statistics to governing bodies and other policymakers.
Conclusions: Rapidly developed, agile extendable LIS functionality and its meaningful use alleviates the administrative burden on laboratory personnel and improves turnaround time of SARS-CoV-2 testing. It will be important to maintain an environment that is conducive for the rapid adoption of meaningful LIS tools after the COVID-19 crisis.
Keywords: COVID-19, laboratory information system, health information technology implementation, computerized provider order entry, change management
Introduction
Background and significance
The novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus has caused a pandemic with unprecedented medical and socioeconomic adversity. Second- and third-wave outbreaks are becoming a reality in several countries, at least partially, due to low herd immunity.[1] Laboratory testing to rapidly detect SARS-CoV-2 using polymerase chain reaction (PCR) is essential to guide proper patient management[2], while serological assays will soon be required to assess population immunity (e.g., antibody tests) and guide national coronavirus disease 2019 (COVID-19) pandemic policies.[1]
The increased demand for round-the-clock laboratory testing during the COVID-19 pandemic has surpassed surge capacity in many clinical laboratories, resulting in significant strain on laboratory personnel and infrastructure.[3][4] These developments are especially undesirable when laboratories already have to process a large numbers of potentially biohazardous specimens every day. Additional delays in testing directly translate into delayed clinical decision making and consequently congest emergency departments and isolation units.[3] Therefore, leveraging the capabilities of laboratory information systems (LIS) to streamline all phases of laboratory testing workflow (preanalytical, analytical, and postanalytical) has proven essential.
To our knowledge, only a single report has been published that discusses the implementation of health informatics to support clinical management of the COVID-19 pandemic through novel electronic health record (EHR) functionality.[5] COVID-19 is a laboratory diagnosis with immediate consequences in the health sector (hospitalization, patient isolation, postponing surgery, etc.). Therefore, clinical laboratories face specific challenges that require dedicated LIS functionality to ensure safe, reliable testing and acceptable turnaround times.[3] However, no literature exists on tools that leverage the LIS in order to alleviate the burden on laboratory personnel, streamline laboratory testing, improve test reporting, facilitate epidemiological and translational research, and enable data-driven policy making.
Objectives
Here, we describe the challenges faced by the Belgian National Reference Center for COVID-19 testing when demand passed allocated surge capacity during the initial phases of the COVID-19 pandemic. We used Kotter’s principles as a framework to rapidly develop and implement additional LIS functionality (Table 1).[6] We implemented tools to manage specimen and data streams, and detail functionality to improve (1) the prelaboratory phase (test ordering, specimen packaging, and shipping), (2) the preanalytical phase (specimen registration, tracking, and test prioritization [triaging]), and (3) the postanalytical phase (automated reporting and facilitating data-driven policymaking). We also briefly discuss unexpected opportunities in which the COVID-19 crisis accelerated the adoption of practices that promote the meaningful use of LIS systems.
| ||||||||||||||||||||||||
Materials and methods
The University Hospitals Leuven is a 2000-bed hospital providing nationwide services via approximately 700,000 consultations, 55,000 admissions, 60,000 emergencies, and 55,000 surgeries annually. The hospital uses a fully in-house–developed EHR that is also commercially available[7] and used in approximately 50% of hospitals in the Flemish Region of Belgium. The clinical laboratory department annually performs around 12,000,000 tests and has national reference functions for several infectious diseases, including the respiratory disease COVID-19. The laboratory performed a total of >25,000 SARS-CoV-2 PCR tests on respiratory samples between February 1 and April 20, 2020. Samples were sent to the reference laboratory from across Belgium, and once analyzed, results were reported to referring clinical laboratories as well as to hospitals and primary care physicians. The LIS is developed in-house and maintained by a dedicated team of computer science engineers and implementation staff. Our LIS includes a computerized physician order entry (CPOE) module for in-house test ordering, which is fully integrated into the EHR.
The LIS automatically sends all validated results to a national EHR database that is directly connected to patient-accessible web-based and mobile applications. All external orders, including those for reference testing, are paper-based and require that request forms accompany the sample. LIS development is directed by a clinical pathologist, allowing for rapid signaling of practical bottlenecks and guidance on the development of health information technology (IT) tools.
Results
COVID-19–specific challenges to laboratory management
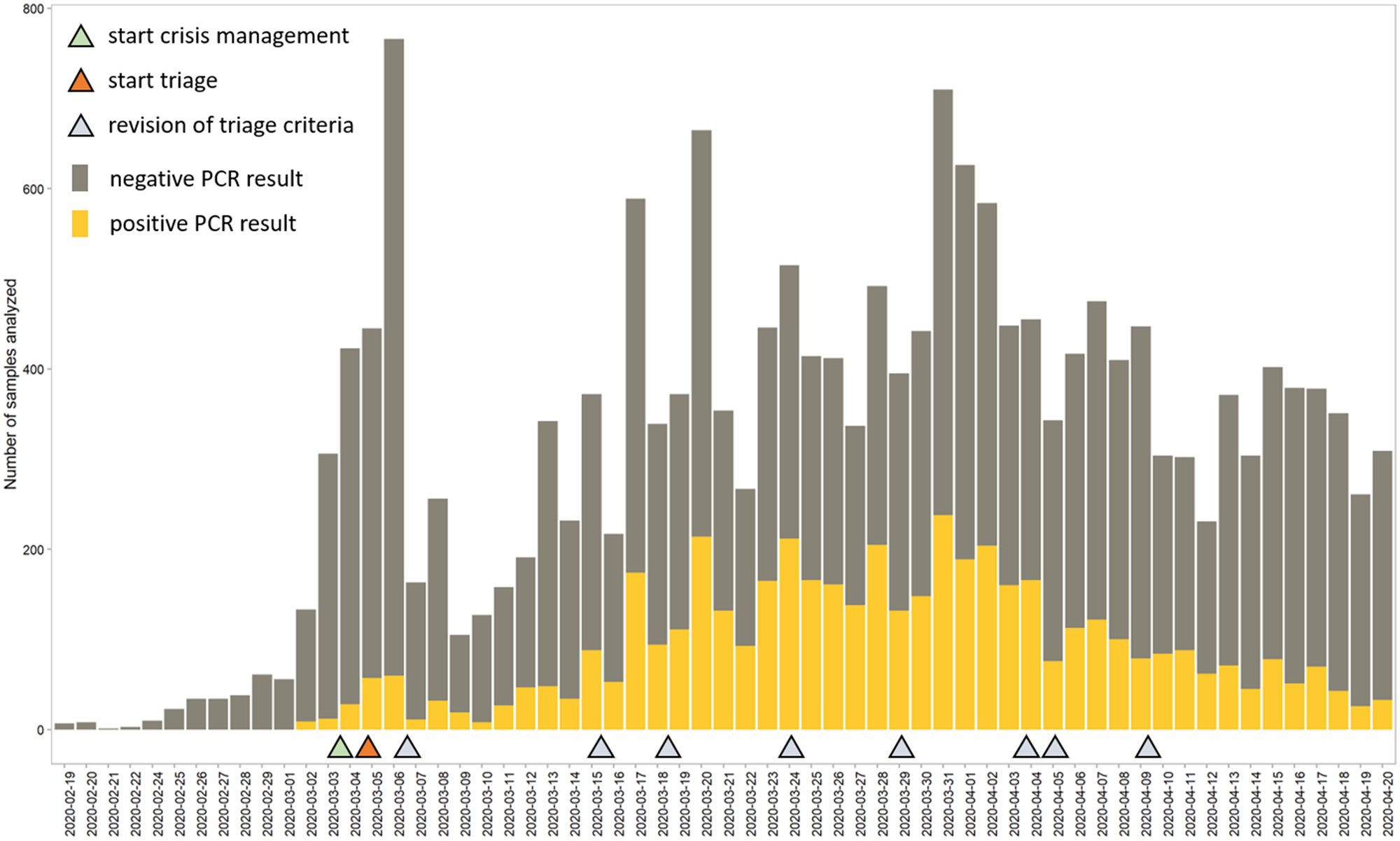
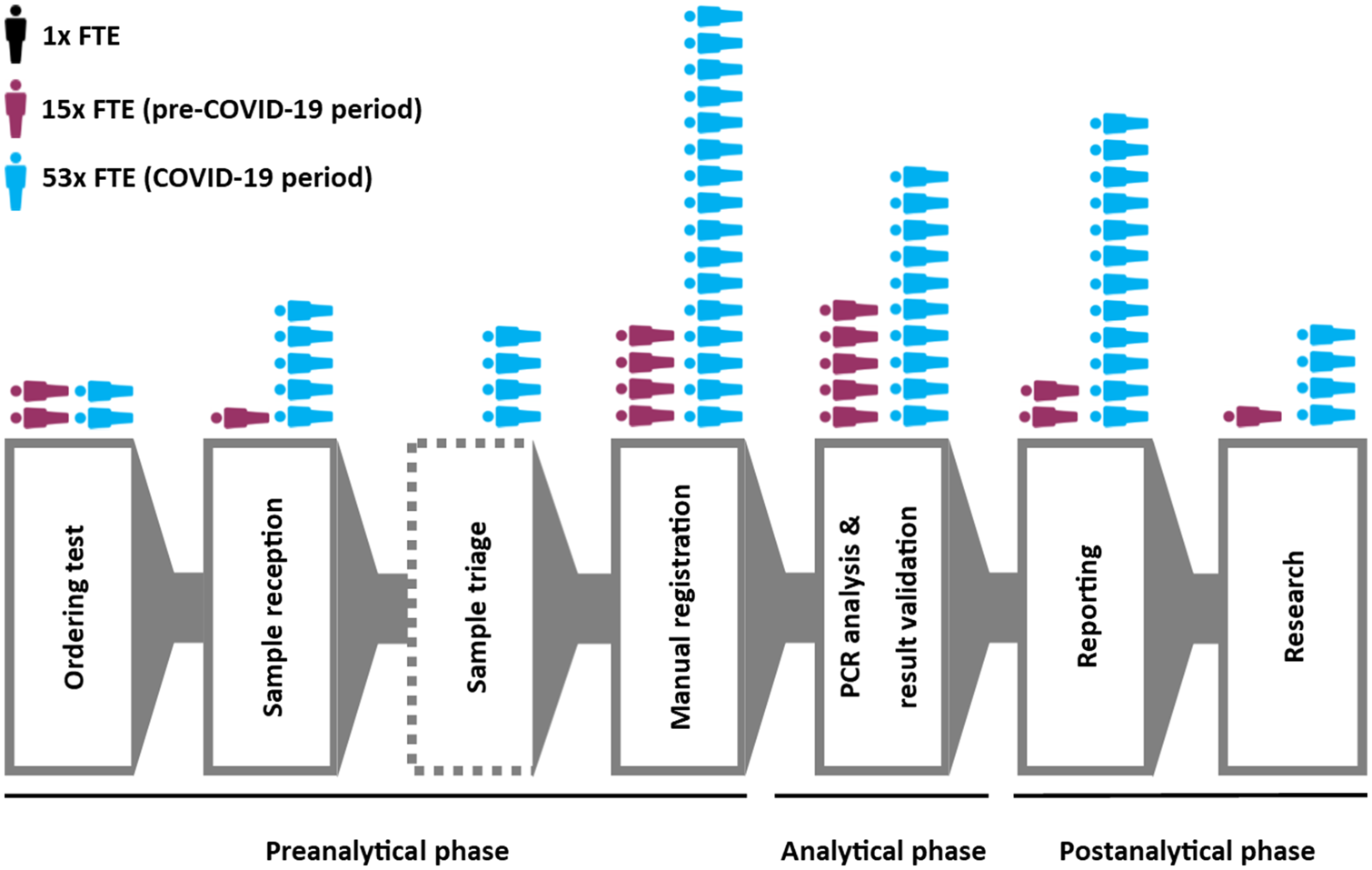
During the early stages of the COVID-19 outbreak, our laboratory was the only SARS-CoV-2 testing center in Belgium. The first case of COVID-19 in Belgium was confirmed by our laboratory on February 3, 2020. The following month, SARS-CoV-2 PCR testing grew exponentially to >750 tests per day, at which point both traditional analytical infrastructure and reagents became limiting factors (Figure 1). To manage the acute crisis, we established a crisis management team (CMT) consisting of clinical pathology staff specialized in microbiology and in LIS development. The CMT readily implemented a test prioritization (triage) system to reduce stress on reagents and addressed key bottlenecks in the preanalytical and postanalytical phase. Maximum upscaling in terms of full-time equivalents (FTEs) was estimated to be an additional 20 FTEs for the preanalytical phase, five for the analytical phase, and 13 for the postanalytical phase (Figure 2). These FTEs were divided among three different shifts. After stabilization of the sample flow, a fraction of laboratory staff was reallocated to perform test validation, leaving some scientific staff to be reallocated to epidemiological studies.
|
|
Notably, the large majority of our expanded workforce (30 of the 38 additional FTEs) was assigned to help with administrative tasks (sample reception, triaging, patient registration, result validation and reporting, and epidemiological studies), and was not directly involved in expanding analytical capacity (i.e., PCR analysis) (Figure 2). By April 2020, reagent supply and machine capacity were more adequate (Figure 1), but pre- and postanalytical bottlenecks could only be resolved by the implementation of dedicated perianalytical IT solutions that reduced administrative burden on laboratory staff by streamlining data flows (Table 2).
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
References
- ↑ 1.0 1.1 Hellewell, J.; Abbott, S.; Gimma, A. et al. (2020). "Feasibility of controlling COVID-19 outbreaks by isolation of cases and contacts". Lancet Global Health 8 (4): e488-e496. doi:10.1016/S2214-109X(20)30074-7. PMC PMC7097845. PMID 32119825. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7097845.
- ↑ Guan, W.-J.; Ni, Z.-Y.; Hu, Y. et al. (2020). "Clinical Characteristics of Coronavirus Disease 2019 in China". New England Journal of Medicine 382 (18): 1708–20. doi:10.1056/NEJMoa2002032. PMC PMC7092819. PMID 32109013. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7092819.
- ↑ 3.0 3.1 3.2 Lippi, G.; Plebani, M. (2020). "The critical role of laboratory medicine during coronavirus disease 2019 (COVID-19) and other viral outbreaks". Clinical Chemistry and Laboratory Medicine 58 (7): 1063–69. doi:10.1515/cclm-2020-0240. PMID 32191623.
- ↑ Posteraro, B.; Marchetti, S.; Roman, L. et al. (2020). "Clinical microbiology laboratory adaptation to COVID-19 emergency: Experience at a large teaching hospital in Rome, Italy". Clinical Microbiology and Infection 26 (8): 1109-1111. doi:10.1016/j.cmi.2020.04.016. PMC 32330569. PMID 32330569. https://www.ncbi.nlm.nih.gov/pmc/articles/32330569.
- ↑ Reeves, J.J.; Hollandsworth, H.M.; Torriani, F.J. et al. (2020). "Rapid response to COVID-19: Health informatics support for outbreak management in an academic health system". Journal of the American Medical Informatics Association 27 (6): 853-859. doi:10.1093/jamia/ocaa037. PMC PMC7184393. PMID 32208481. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7184393.
- ↑ 6.0 6.1 Kotter, J.P. (May–June 1995). "Leading Change: Why Transformation Efforts Fail". Harvard Business Review. https://hbr.org/1995/05/leading-change-why-transformation-efforts-fail-2. Retrieved 22 April 2020.
- ↑ "NexuzHealth". Nexuzhealth. https://www.nexuzhealth.be/en. Retrieved 22 April 2020.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added.