Difference between revisions of "Journal:Intervene: A tool for intersection and visualization of multiple gene or genomic region sets"
Shawndouglas (talk | contribs) (Created stub. Saving and adding more.) |
Shawndouglas (talk | contribs) (Saving and adding more.) |
||
| Line 36: | Line 36: | ||
There are several web applications and [[R (programming language)|R packages] available to compute intersection and visualization of up to six list sets by using Venn diagrams. Although tools exist to perform genomic region set intersections<ref name="ZhuChIP10">{{cite journal |title=ChIPpeakAnno: A Bioconductor package to annotate ChIP-seq and ChIP-chip data |journal=BMC Bioinformatics |author=Zhu, L.J.; Gazin, C.; Lawson, N.D. et al. |volume=11 |pages=237 |year=2010 |doi=10.1186/1471-2105-11-237 |pmid=20459804 |pmc=PMC3098059}}</ref><ref name="DalePyebedtools11">{{cite journal |title=Pybedtools: A flexible Python library for manipulating genomic datasets and annotations |journal=Bioinformatics |author=Dale, R.K.; Pedersen, B.S.; Quinlan, A.R. |volume=27 |issue=24 |pages=3423–4 |year=2011 |doi=10.1093/bioinformatics/btr539 |pmid=21949271 |pmc=PMC3232365}}</ref><ref name="HunterMatplotlib07">{{cite journal |title=Matplotlib: A 2D Graphics Environment |journal=Computing in Science & Engineering |author=Hunter, J.D. |volume=9 |issue=3 |year=2007 |doi=10.1109/MCSE.2007.55}}</ref>, there is a limited number of tools available to visualize them.<ref name="ZhuChIP10" /><ref name="DalePyebedtools11" /> To our knowledge no tool exists to generate UpSet plots for genomic region sets. Consequently, there is a great need for integrative tools to compute and visualize intersection of multiple sets of both genomic regions and gene/list sets. | There are several web applications and [[R (programming language)|R packages] available to compute intersection and visualization of up to six list sets by using Venn diagrams. Although tools exist to perform genomic region set intersections<ref name="ZhuChIP10">{{cite journal |title=ChIPpeakAnno: A Bioconductor package to annotate ChIP-seq and ChIP-chip data |journal=BMC Bioinformatics |author=Zhu, L.J.; Gazin, C.; Lawson, N.D. et al. |volume=11 |pages=237 |year=2010 |doi=10.1186/1471-2105-11-237 |pmid=20459804 |pmc=PMC3098059}}</ref><ref name="DalePyebedtools11">{{cite journal |title=Pybedtools: A flexible Python library for manipulating genomic datasets and annotations |journal=Bioinformatics |author=Dale, R.K.; Pedersen, B.S.; Quinlan, A.R. |volume=27 |issue=24 |pages=3423–4 |year=2011 |doi=10.1093/bioinformatics/btr539 |pmid=21949271 |pmc=PMC3232365}}</ref><ref name="HunterMatplotlib07">{{cite journal |title=Matplotlib: A 2D Graphics Environment |journal=Computing in Science & Engineering |author=Hunter, J.D. |volume=9 |issue=3 |year=2007 |doi=10.1109/MCSE.2007.55}}</ref>, there is a limited number of tools available to visualize them.<ref name="ZhuChIP10" /><ref name="DalePyebedtools11" /> To our knowledge no tool exists to generate UpSet plots for genomic region sets. Consequently, there is a great need for integrative tools to compute and visualize intersection of multiple sets of both genomic regions and gene/list sets. | ||
To address this need, we developed Intervene, an easy-to-use command line tool to compute and visualize intersections of genomic regions with Venn diagrams, UpSet plots, or clustered heat maps. Moreover, we provide an interactive web application companion to upload list sets or the output of Intervene to further customize plots. | To address this need, we developed Intervene, an easy-to-use command line tool to compute and visualize intersections of genomic regions with Venn diagrams, UpSet plots, or clustered heat maps. Moreover, we provide an interactive web application companion to upload list sets or the output of Intervene to further customize plots. | ||
==Implementation== | |||
Intervene comes as a command line tool, along with an interactive Shiny web application to customize the visual representation of intersections. The command line tool is implemented in Python (version 2.7) and R programming language (version 3.3.2). The build also works with Python versions 3.4, 3.5, and 3.6. The accompanying web interface is developed using Shiny (version 1.0.0), a web application framework for R. Intervene uses pybedtools<ref name="DalePyebedtools11" /> to perform genomic region set intersections and Seaborn (https://seaborn.pydata.org/), Matplotlib<ref name="HunterMatplotlib07" />, UpSetR<ref name="ConwayUpSetR17">{{cite journal |title=UpSetR: An R Package For The Visualization Of Intersecting Sets And Their Properties |journal=bioRxiv |author=Conway, J.R.; Lex, A.; Gehlenborg, N. |date=25 March 2017 |doi=10.1101/120600}}</ref>, and Corrplot<ref name="WeiCorrplot16">{{cite web |url=https://cran.r-project.org/package=corrplot |title=corrplot: Visualization of a Correlation Matrix |author=Wei, T.; Simko, V. |date=21 April 2016}}</ref> to generate figures. The web application uses the R package Venerable<ref name="SwintonVenn09">{{cite web |url=https://r-forge.r-project.org/scm/viewvc.php/*checkout*/pkg/Vennerable/inst/doc/Venn.pdf?revision=58&root=vennerable |title=Venn diagrams in R with Vennerable package |author=Swinton, J. |date=23 September 2009}}</ref> for different types of Venn diagrams, UpSetR for UpSet plots, and heatmap.2 and Corrplot for pairwise intersection clustered heat maps. The UpSet module of the web ShinyApp was derived from the UpSetR<ref name="ConwayUpSetR17" /> ShinyApp, which was extended by adding more options and features to customize the UpSet plots. | |||
Intervene can be installed by using ''pip install intervene'' or using the source code available on bitbucket https://bitbucket.org/CBGR/intervene. The tool has been tested on Linux and MAC systems. The Shiny web application is hosted with shinyapps.io by RStudio, and is compatible with all modern web browsers. A detailed documentation including installation instructions and how to use the tool is provided in Additional file 1 and is available at http://intervene.readthedocs.io. | |||
==Results== | |||
===An integrated tool for effective visualization of multiple set intersections=== | |||
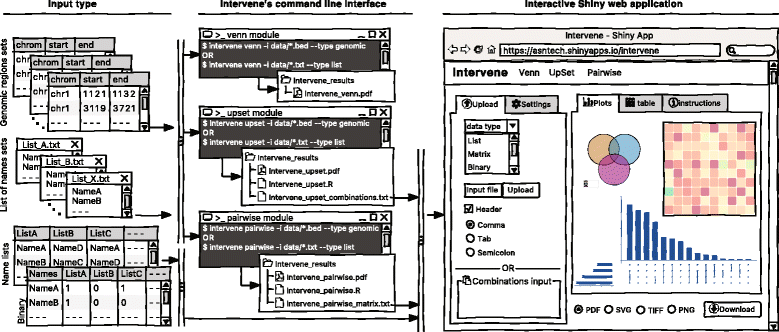
As visualization of sets and their intersections is becoming more and more challenging due to the increasing number of generated data sets, there is a strong need to have an integrated tool to compute and visualize intersections effectively. To address this challenge, we have developed Intervene, which is composed of three different modules, accessible through the subcommands ''venn'', ''upset'', and ''pairwise''. Intervene accepts two types of input files: genomic regions in BED, GFF, or VCF format and gene/name lists in plain text format. A detailed sketch of Intervene’s command line interface and web application utility with types of inputs is provided in Fig. 1. | |||
[[File:Fig1 Khan BMCBioinformatics2017 18.gif|779px]] | |||
{{clear}} | |||
{| | |||
| STYLE="vertical-align:top;"| | |||
{| border="0" cellpadding="5" cellspacing="0" width="779px" | |||
|- | |||
| style="background-color:white; padding-left:10px; padding-right:10px;"| <blockquote>'''Figure 1.''' A sketch of Intervene’s command line interface and web application, and input data type</blockquote> | |||
|- | |||
|} | |||
|} | |||
Intervene provides flexibility to the user to choose figure colors, label text, size, resolution, and type to make them publication-standard quality. To read the help about any module, the user can type ''intervene < subcommand > −-help'' on the command line. Furthermore, Intervene produces results as text files, which can be easily imported to the web application for interactive visualization and customization of plots (see “An interactive web application” section). | |||
==References== | ==References== | ||
Revision as of 01:14, 6 June 2017
| Full article title | Intervene: A tool for intersection and visualization of multiple gene or genomic region sets |
|---|---|
| Journal | BMC Bioinformatics |
| Author(s) | Khan, Aziz; Mathelier, Anthony |
| Author affiliation(s) | Centre for Molecular Medicine Norway, Norwegian Radium Hospital |
| Primary contact | Email: aziz dot khan at ncmm dot uio dot no and anthony dot mathelier at ncmm dot uio dot no |
| Year published | 2017 |
| Volume and issue | 18 |
| Page(s) | 287 |
| DOI | 10.1186/s12859-017-1708-7 |
| ISSN | 1471-2105 |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-017-1708-7 |
| Download | https://bmcbioinformatics.biomedcentral.com/track/pdf/10.1186/s12859-017-1708-7 (PDF) |
Abstract
Background: A common task for scientists relies on comparing lists of genes or genomic regions derived from high-throughput sequencing experiments. While several tools exist to intersect and visualize sets of genes, similar tools dedicated to the visualization of genomic region sets are currently limited.
Results: To address this gap, we have developed the Intervene tool, which provides an easy and automated interface for the effective intersection and visualization of genomic region or list sets, thus facilitating their analysis and interpretation. Intervene contains three modules: venn to generate Venn diagrams of up to six sets, upset to generate UpSet plots of multiple sets, and pairwise to compute and visualize intersections of multiple sets as clustered heat maps. Intervene, and its interactive web ShinyApp companion, generate publication-quality figures for the interpretation of genomic region and list sets.
Conclusions: Intervene and its web application companion provide an easy command line and an interactive web interface to compute intersections of multiple genomic and list sets. They have the capacity to plot intersections using easy-to-interpret visual approaches. Intervene is developed and designed to meet the needs of both computer scientists and biologists. The source code is freely available at https://bitbucket.org/CBGR/intervene, with the web application available at https://asntech.shinyapps.io/intervene.
Keywords: visualization, Venn diagrams, UpSet plots, heat maps, genome analysis
Background
Effective visualization of transcriptomic, genomic, and epigenomic data generated by next-generation sequencing-based high-throughput assays have become an area of great interest. Most of the data sets generated by such assays are lists of genes or variants, and genomic region sets. The genomic region sets represent genomic locations for specific features, such as transcription factor – DNA interactions, transcription start sites, histone modifications, and DNase hypersensitivity sites. A common task in the interpretation of these features is to find similarities, differences, and enrichments between such sets, which come from different samples, experimental conditions, or cell and tissue types.
Classically, the intersection or overlap between different sets, such as gene lists, is represented by Venn diagrams[1] or Edwards-Venn.[2] If the number of sets exceeds four, such diagrams become complex and difficult to interpret. The key challenge is that there are 2n combinations to visually represent when considering n sets. An alternative approach, the UpSet plots, was introduced to depict the intersection of more than three sets.[3] The advantage of UpSet plots is their capacity to rank the intersections and alternatively hide combinations without intersection, which is not possible using a Venn diagram. However, with a large number of sets, UpSet plots become an ineffective way of illustrating set intersections. To visualize a large number of sets, one can represent pairwise intersections using a clustered heat map as suggested by Lex and Gehlenborg.[4]
There are several web applications and [[R (programming language)|R packages] available to compute intersection and visualization of up to six list sets by using Venn diagrams. Although tools exist to perform genomic region set intersections[5][6][7], there is a limited number of tools available to visualize them.[5][6] To our knowledge no tool exists to generate UpSet plots for genomic region sets. Consequently, there is a great need for integrative tools to compute and visualize intersection of multiple sets of both genomic regions and gene/list sets.
To address this need, we developed Intervene, an easy-to-use command line tool to compute and visualize intersections of genomic regions with Venn diagrams, UpSet plots, or clustered heat maps. Moreover, we provide an interactive web application companion to upload list sets or the output of Intervene to further customize plots.
Implementation
Intervene comes as a command line tool, along with an interactive Shiny web application to customize the visual representation of intersections. The command line tool is implemented in Python (version 2.7) and R programming language (version 3.3.2). The build also works with Python versions 3.4, 3.5, and 3.6. The accompanying web interface is developed using Shiny (version 1.0.0), a web application framework for R. Intervene uses pybedtools[6] to perform genomic region set intersections and Seaborn (https://seaborn.pydata.org/), Matplotlib[7], UpSetR[8], and Corrplot[9] to generate figures. The web application uses the R package Venerable[10] for different types of Venn diagrams, UpSetR for UpSet plots, and heatmap.2 and Corrplot for pairwise intersection clustered heat maps. The UpSet module of the web ShinyApp was derived from the UpSetR[8] ShinyApp, which was extended by adding more options and features to customize the UpSet plots.
Intervene can be installed by using pip install intervene or using the source code available on bitbucket https://bitbucket.org/CBGR/intervene. The tool has been tested on Linux and MAC systems. The Shiny web application is hosted with shinyapps.io by RStudio, and is compatible with all modern web browsers. A detailed documentation including installation instructions and how to use the tool is provided in Additional file 1 and is available at http://intervene.readthedocs.io.
Results
An integrated tool for effective visualization of multiple set intersections
As visualization of sets and their intersections is becoming more and more challenging due to the increasing number of generated data sets, there is a strong need to have an integrated tool to compute and visualize intersections effectively. To address this challenge, we have developed Intervene, which is composed of three different modules, accessible through the subcommands venn, upset, and pairwise. Intervene accepts two types of input files: genomic regions in BED, GFF, or VCF format and gene/name lists in plain text format. A detailed sketch of Intervene’s command line interface and web application utility with types of inputs is provided in Fig. 1.
|
Intervene provides flexibility to the user to choose figure colors, label text, size, resolution, and type to make them publication-standard quality. To read the help about any module, the user can type intervene < subcommand > −-help on the command line. Furthermore, Intervene produces results as text files, which can be easily imported to the web application for interactive visualization and customization of plots (see “An interactive web application” section).
References
- ↑ Venn, J. (1880). "On the diagrammatic and mechanical representation of propositions and reasonings". Philisophical Magazine and Journal of Science 10 (59): 1–18. doi:10.1080/14786448008626877.
- ↑ Edwards, A.W.F. (2004). Cogwheels of the Mind: The Story of Venn Diagrams. Johns Hopkins University Press. pp. 128. ISBN 9780801874345.
- ↑ Lex, A.; Gehlenborg, N.; Strobelt, H. et al. (2014). "UpSet: Visualization of Intersecting Sets". IEEE Transactions on Visualization and Computer Graphics 20 (12): 1983-92. doi:10.1109/TVCG.2014.2346248.
- ↑ Lex, A.; Gehlenborg, N. (2014). "Points of view: Sets and intersections". IEEE Transactions on Visualization and Computer Graphics 11: 779. doi:10.1038/nmeth.3033.
- ↑ 5.0 5.1 Zhu, L.J.; Gazin, C.; Lawson, N.D. et al. (2010). "ChIPpeakAnno: A Bioconductor package to annotate ChIP-seq and ChIP-chip data". BMC Bioinformatics 11: 237. doi:10.1186/1471-2105-11-237. PMC PMC3098059. PMID 20459804. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3098059.
- ↑ 6.0 6.1 6.2 Dale, R.K.; Pedersen, B.S.; Quinlan, A.R. (2011). "Pybedtools: A flexible Python library for manipulating genomic datasets and annotations". Bioinformatics 27 (24): 3423–4. doi:10.1093/bioinformatics/btr539. PMC PMC3232365. PMID 21949271. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3232365.
- ↑ 7.0 7.1 Hunter, J.D. (2007). "Matplotlib: A 2D Graphics Environment". Computing in Science & Engineering 9 (3). doi:10.1109/MCSE.2007.55.
- ↑ 8.0 8.1 Conway, J.R.; Lex, A.; Gehlenborg, N. (25 March 2017). "UpSetR: An R Package For The Visualization Of Intersecting Sets And Their Properties". bioRxiv. doi:10.1101/120600.
- ↑ Wei, T.; Simko, V. (21 April 2016). "corrplot: Visualization of a Correlation Matrix". https://cran.r-project.org/package=corrplot.
- ↑ Swinton, J. (23 September 2009). "Venn diagrams in R with Vennerable package". https://r-forge.r-project.org/scm/viewvc.php/*checkout*/pkg/Vennerable/inst/doc/Venn.pdf?revision=58&root=vennerable.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. In some cases important information was missing from the references, and that information was added. Some grammar were corrected when necessary.