Difference between revisions of "Main Page/Featured article of the week/2018"
Shawndouglas (talk | contribs) (Added last week's article of the week.) |
Shawndouglas (talk | contribs) (Added last week's article of the week.) |
||
| Line 17: | Line 17: | ||
<!-- Below this line begin pasting previous news --> | <!-- Below this line begin pasting previous news --> | ||
<h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: February 19–25:</h2> | <h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: February 26–March 4:</h2> | ||
<div style="float: left; margin: 0.5em 0.9em 0.4em 0em;">[[File:Fig4 Scotti Molecules2018 23-1.png|240px]]</div> | |||
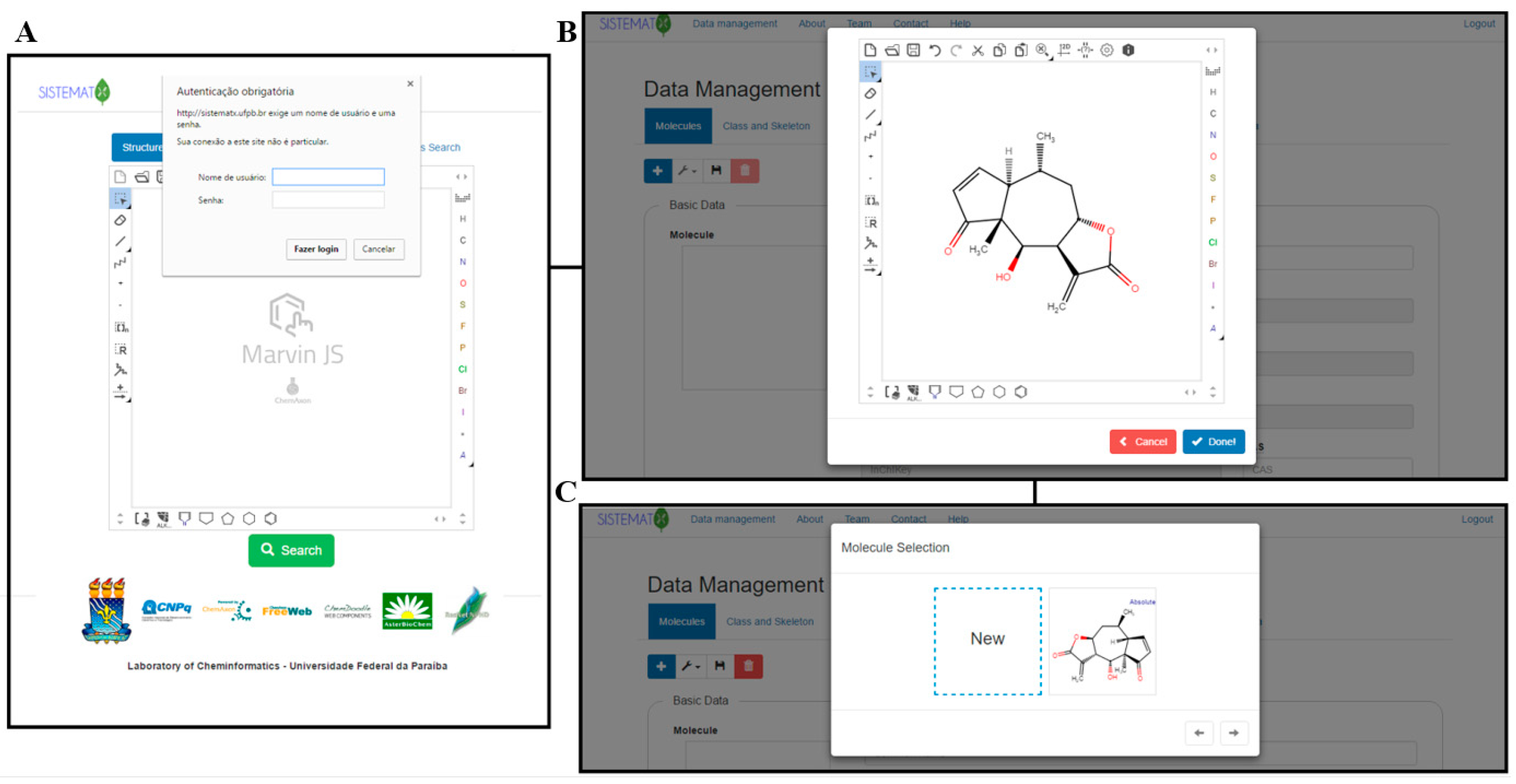
'''"[[Journal:SistematX, an online web-based cheminformatics tool for data management of secondary metabolites|SistematX, an online web-based cheminformatics tool for data management of secondary metabolites]]"''' | |||
The traditional work of a natural products researcher consists in large part of time-consuming experimental work, collecting biota to prepare, extracts to analyze, and innovative metabolites to identify. However, along this long scientific path, much information is lost or restricted to a specific niche. The large amounts of data already produced and the science of metabolomics reveal new questions: Are these compounds known or new? How fast can this information be obtained? To answer these and other relevant questions, an appropriate procedure to correctly store [[information]] on the data retrieved from the discovered metabolites is necessary. The SistematX (http://sistematx.ufpb.br) interface is implemented considering the following aspects: (a) the ability to search by structure, [[SMILES (language)|SMILES]] (Simplified Molecular-Input Line-Entry System) code, compound name, and species; (b) the ability to save chemical structures found by searching; (c) the ability to display compound data results, including important characteristics for natural products chemistry; and (d) the user's ability to find specific information for taxonomic rank (from family to species) of the plant from which the compound was isolated, the searched-for molecule, and the bibliographic reference and Global Positioning System (GPS) coordinates. ('''[[Journal:SistematX, an online web-based cheminformatics tool for data management of secondary metabolites|Full article...]]''')<br /> | |||
|- | |||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: February 19–25:</h2> | |||
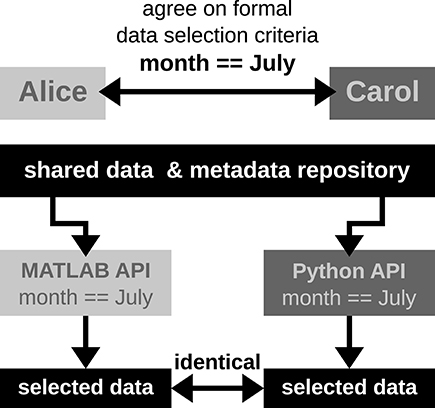
'''"[[Journal:Rethinking data sharing and human participant protection in social science research: Applications from the qualitative realm|Rethinking data sharing and human participant protection in social science research: Applications from the qualitative realm]]"''' | '''"[[Journal:Rethinking data sharing and human participant protection in social science research: Applications from the qualitative realm|Rethinking data sharing and human participant protection in social science research: Applications from the qualitative realm]]"''' | ||
While data sharing is becoming increasingly common in quantitative social inquiry, qualitative data are rarely shared. One factor inhibiting data sharing is a concern about human participant protections and privacy. Protecting the confidentiality and safety of research participants is a concern for both quantitative and qualitative researchers, but it raises specific concerns within the epistemic context of qualitative research. Thus, the applicability of emerging protection models from the quantitative realm must be carefully evaluated for application to the qualitative realm. At the same time, qualitative scholars already employ a variety of strategies for human-participant protection implicitly or informally during the research process. In this practice paper, we assess available strategies for protecting human participants and how they can be deployed. We describe a spectrum of possible data management options, such as de-identification and applying access controls, including some already employed by the Qualitative Data Repository (QDR) in tandem with its pilot depositors. ('''[[Journal:Rethinking data sharing and human participant protection in social science research: Applications from the qualitative realm|Full article...]]''')<br /> | While data sharing is becoming increasingly common in quantitative social inquiry, qualitative data are rarely shared. One factor inhibiting data sharing is a concern about human participant protections and privacy. Protecting the confidentiality and safety of research participants is a concern for both quantitative and qualitative researchers, but it raises specific concerns within the epistemic context of qualitative research. Thus, the applicability of emerging protection models from the quantitative realm must be carefully evaluated for application to the qualitative realm. At the same time, qualitative scholars already employ a variety of strategies for human-participant protection implicitly or informally during the research process. In this practice paper, we assess available strategies for protecting human participants and how they can be deployed. We describe a spectrum of possible data management options, such as de-identification and applying access controls, including some already employed by the Qualitative Data Repository (QDR) in tandem with its pilot depositors. ('''[[Journal:Rethinking data sharing and human participant protection in social science research: Applications from the qualitative realm|Full article...]]''')<br /> | ||
|- | |- | ||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: February 12–18:</h2> | |<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: February 12–18:</h2> | ||
| Line 28: | Line 33: | ||
To date, non-reproducibility of neurophysiological research is a matter of intense discussion in the scientific community. A crucial component to enhance reproducibility is to comprehensively collect and store metadata, that is, all information about the experiment, the data, and the applied preprocessing steps on the data, such that they can be accessed and shared in a consistent and simple manner. However, the complexity of experiments, the highly specialized analysis workflows, and a lack of knowledge on how to make use of supporting software tools often overburden researchers to perform such a detailed documentation. For this reason, the collected metadata are often incomplete, incomprehensible for outsiders, or ambiguous. Based on our research experience in dealing with diverse datasets, we here provide conceptual and technical guidance to overcome the challenges associated with the collection, organization, and storage of metadata in a neurophysiology [[laboratory]]. ('''[[Journal:Handling metadata in a neurophysiology laboratory|Full article...]]''')<br /> | To date, non-reproducibility of neurophysiological research is a matter of intense discussion in the scientific community. A crucial component to enhance reproducibility is to comprehensively collect and store metadata, that is, all information about the experiment, the data, and the applied preprocessing steps on the data, such that they can be accessed and shared in a consistent and simple manner. However, the complexity of experiments, the highly specialized analysis workflows, and a lack of knowledge on how to make use of supporting software tools often overburden researchers to perform such a detailed documentation. For this reason, the collected metadata are often incomplete, incomprehensible for outsiders, or ambiguous. Based on our research experience in dealing with diverse datasets, we here provide conceptual and technical guidance to overcome the challenges associated with the collection, organization, and storage of metadata in a neurophysiology [[laboratory]]. ('''[[Journal:Handling metadata in a neurophysiology laboratory|Full article...]]''')<br /> | ||
|- | |- | ||
|<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: February 5–11:</h2> | |<br /><h2 style="font-size:105%; font-weight:bold; text-align:left; color:#000; padding:0.2em 0.4em; width:50%;">Featured article of the week: February 5–11:</h2> | ||
Revision as of 21:00, 5 March 2018
|
|
If you're looking for other "Article of the Week" archives: 2014 - 2015 - 2016 - 2017 - 2018 |
Featured article of the week archive - 2018
Welcome to the LIMSwiki 2018 archive for the Featured Article of the Week.
Featured article of the week: February 26–March 4:"SistematX, an online web-based cheminformatics tool for data management of secondary metabolites" The traditional work of a natural products researcher consists in large part of time-consuming experimental work, collecting biota to prepare, extracts to analyze, and innovative metabolites to identify. However, along this long scientific path, much information is lost or restricted to a specific niche. The large amounts of data already produced and the science of metabolomics reveal new questions: Are these compounds known or new? How fast can this information be obtained? To answer these and other relevant questions, an appropriate procedure to correctly store information on the data retrieved from the discovered metabolites is necessary. The SistematX (http://sistematx.ufpb.br) interface is implemented considering the following aspects: (a) the ability to search by structure, SMILES (Simplified Molecular-Input Line-Entry System) code, compound name, and species; (b) the ability to save chemical structures found by searching; (c) the ability to display compound data results, including important characteristics for natural products chemistry; and (d) the user's ability to find specific information for taxonomic rank (from family to species) of the plant from which the compound was isolated, the searched-for molecule, and the bibliographic reference and Global Positioning System (GPS) coordinates. (Full article...)
|